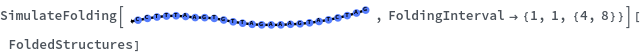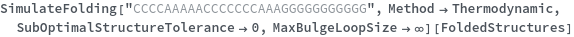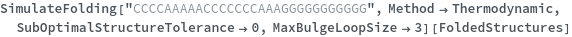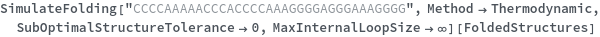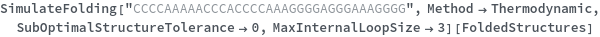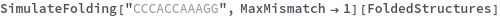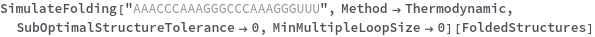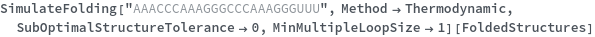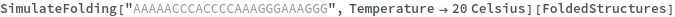SimulateFolding
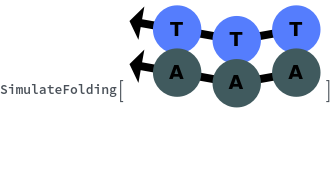
SimulateFolding[oligomer]⟹foldingObject
predicts potential secondary structure interactions of the provided oligomer with itself.
Details
- Kinetic method is an implementation of Nussinov's dynamic programing algorithm described in Object[Report, Literature, "id:E8zoYveXdRWm"]: R. Nussinov et al. "Algorithms for loop matchings." SIAM Journal on Applied mathematics 35.1 (1978): 68-82.
- Thermodynamic method is an implementation of Zuker's free energy minimization algorithm described in Object[Report, Literature, "id:GmzlKjYVz86e"]: M. Zuker. "On finding all suboptimal foldings of an RNA molecule." Science 244.4900 (1989): 48-100.
- The Thermodynamic folding allows only for planar folding and thus can not detect so called "pseudoknots", which can be found by the Kinetic method. However, structures returned by the Kinetic method places no constraints on which bases can pair and therefore may not necessarily be topologically obtainable.
- Following loops can appear in folded structures:
-
LoopDescription
StackingLoop Watson-Crick paired nucleotides on both strands. BulgeLoop Unpaired nucleotides occur on only one strand of double helix. InternalLoop Double helix interrupted by non-Watson-Crick paired nucleotides on both strands. MultipleLoop Three or more helixes intersect.
Input
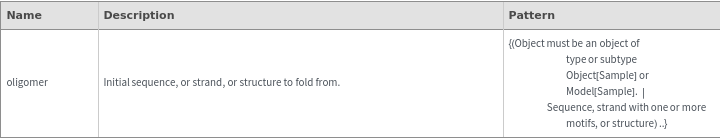
Output
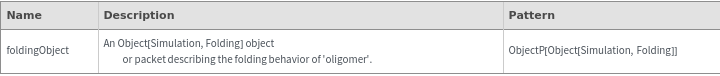
General Options
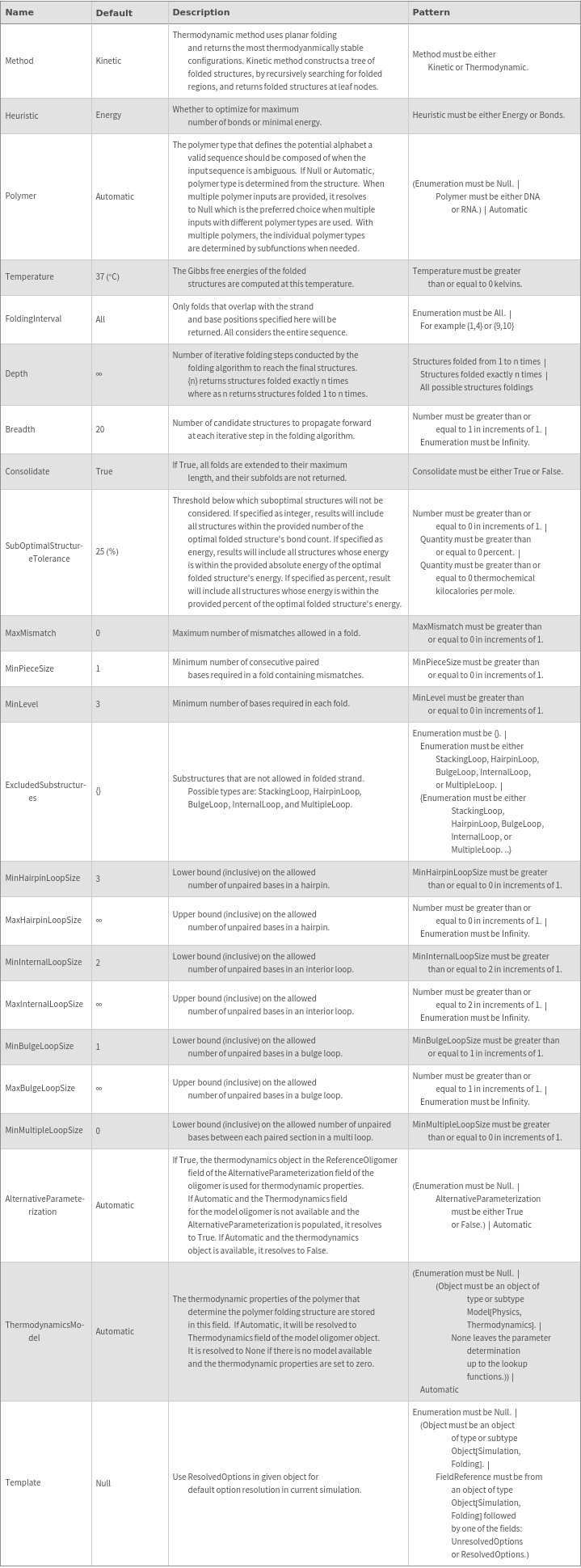
Examples
Basic Examples (5)
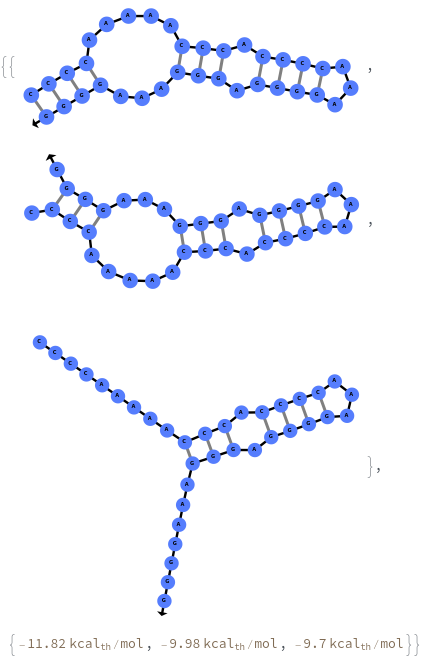
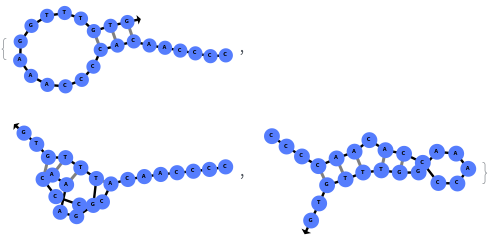
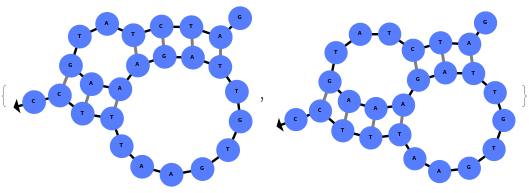
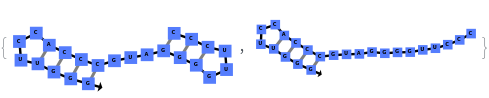
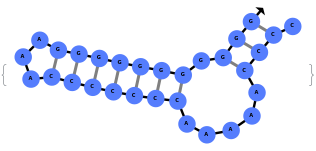
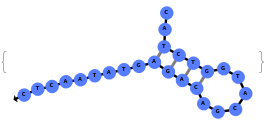
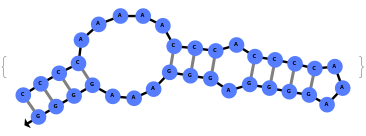
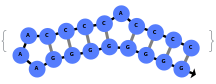
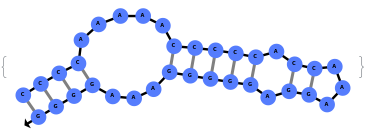
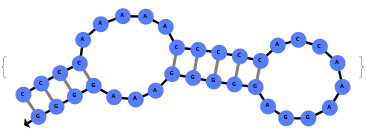
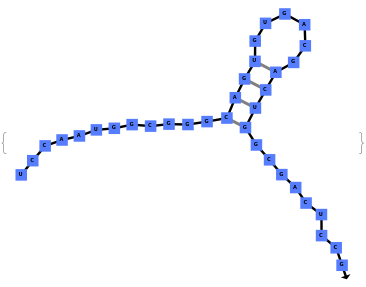
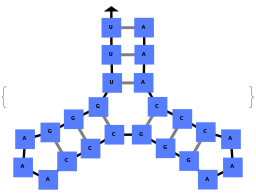
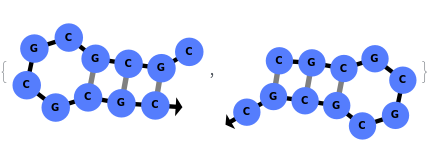
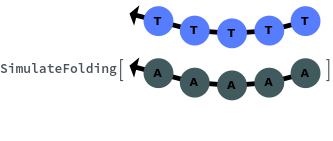
When provided with a sequence, SimulateFolding predicts potentially high energy secondary structures the sequence could fold into through Watson-Crick base pairing rules:

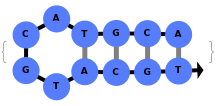
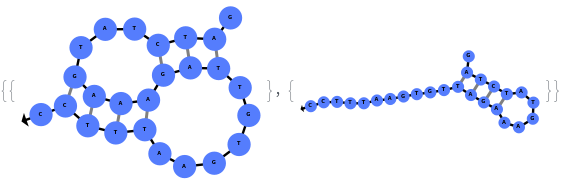
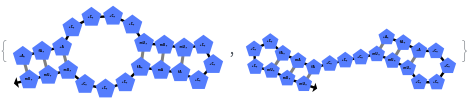
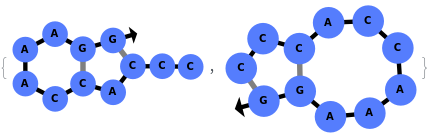
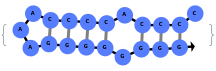
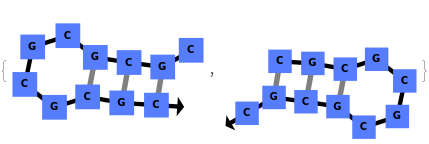
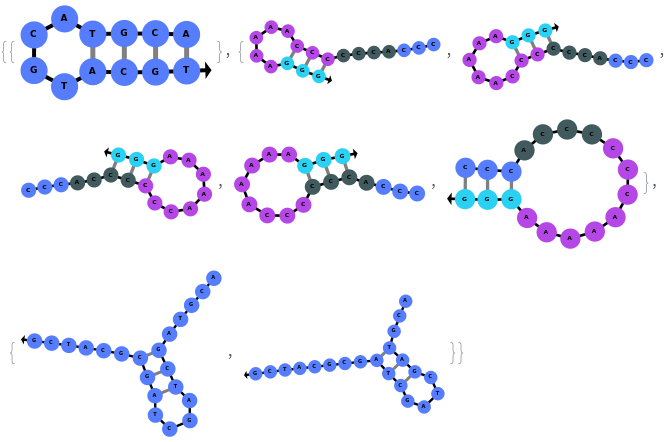
In addition to accepting sequences, the function can handle strands with more than one sequence:

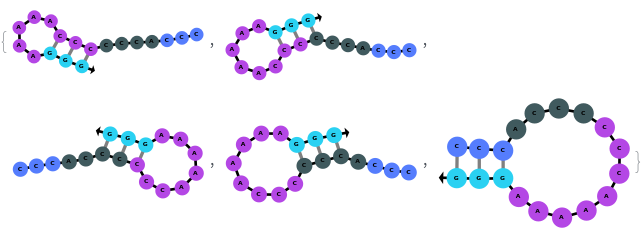
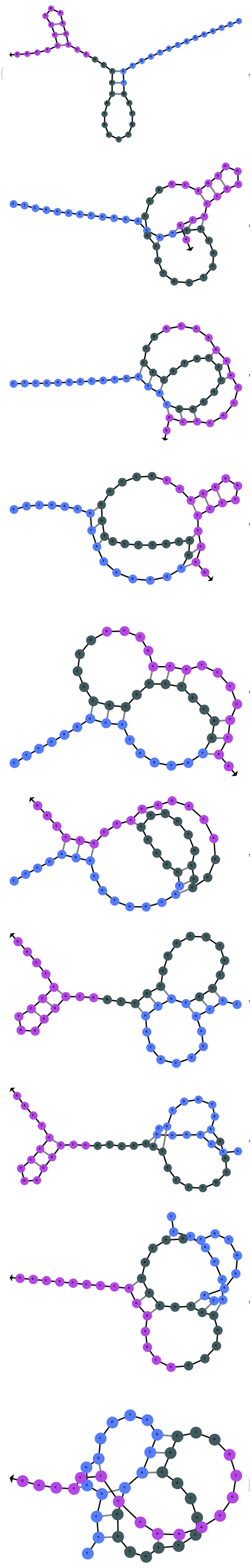
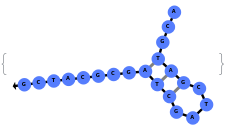
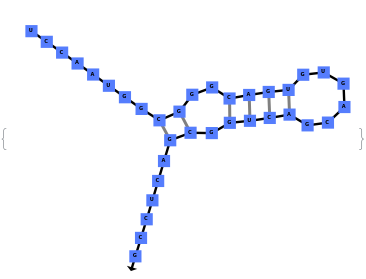
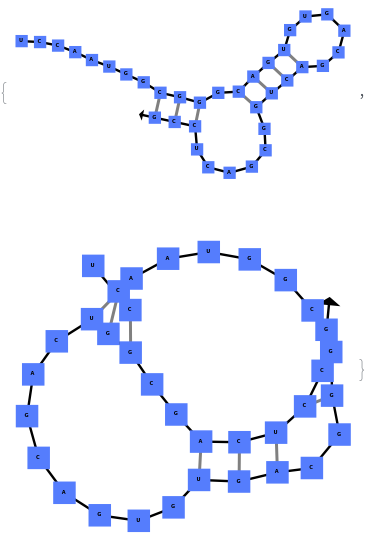
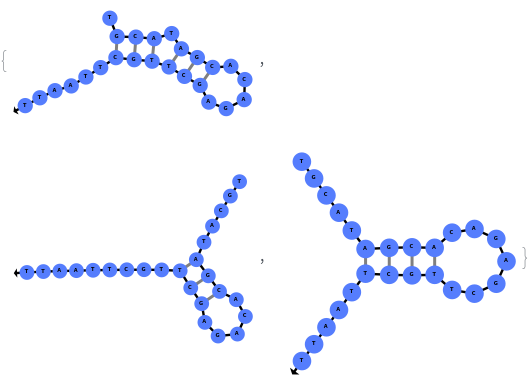
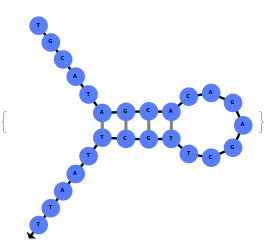
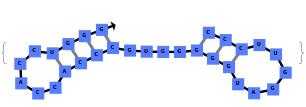
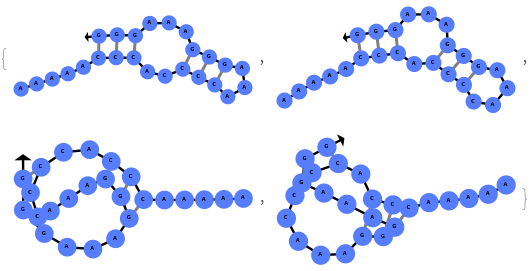
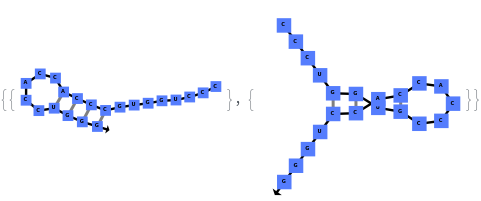
The function can also handled complicate structures where it considers the high energy structures that might further form through folding:
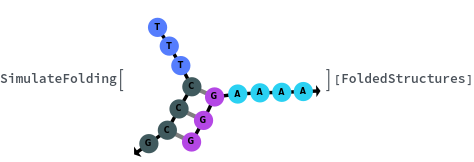
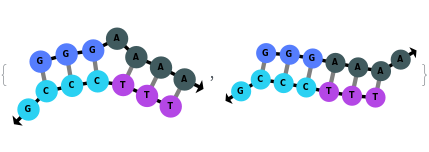
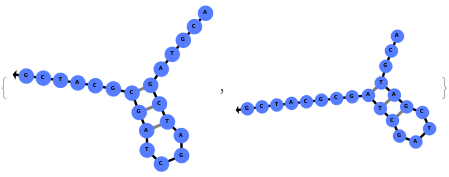
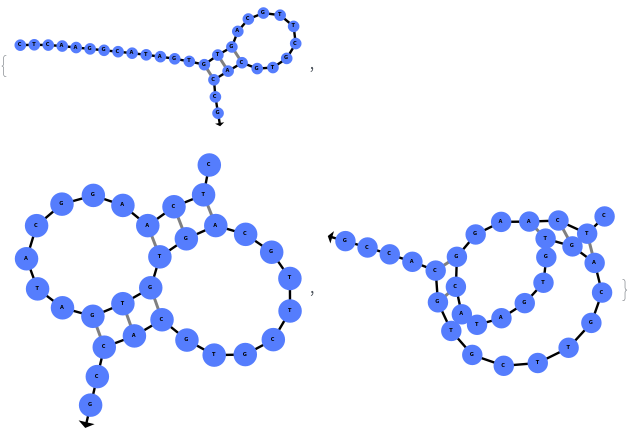
The function can also be applied to oligomer samples:


The function can also be applied to oligomer models:

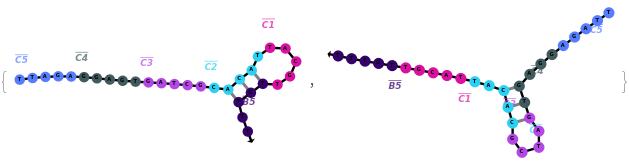
Options (41)
AlternativeParameterization (1)
ExcludedSubstructures (3)
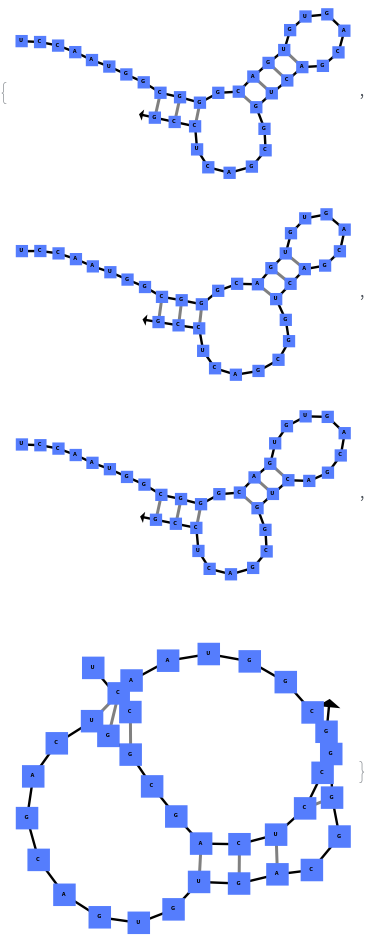
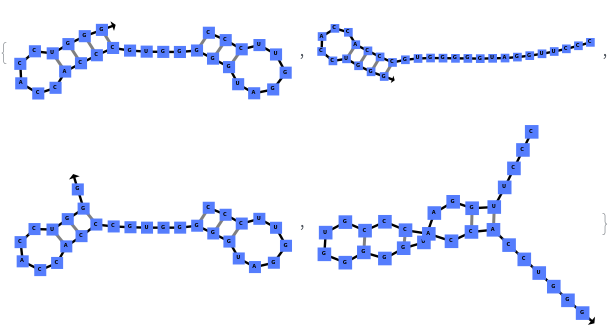
When you exclude no substructures (all are allowed):

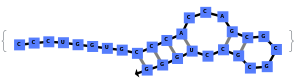
By excluding internal loops (this includes mismatches), we get the most optimal structure that does not have internal loops:

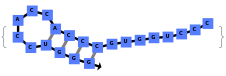
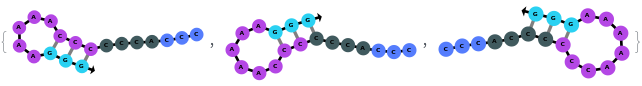
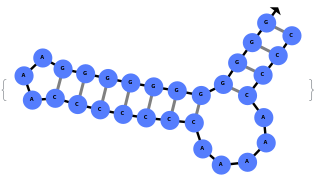
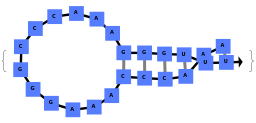
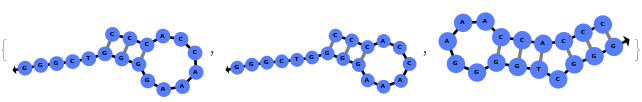
By excluding bulge loops and internal loops, we get the new most optimal structure (no bulge/internal loops) and a structure whose energy is within 25% of the optimal structure's energy: