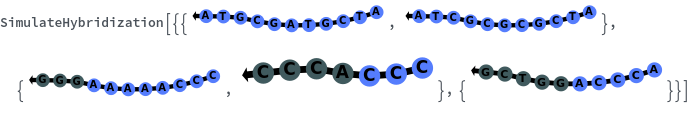
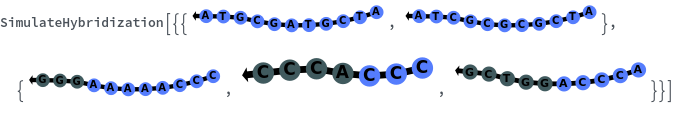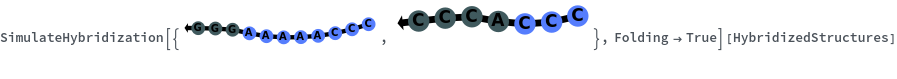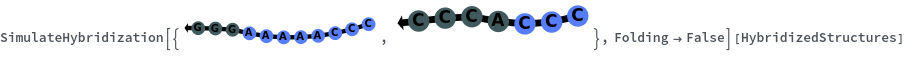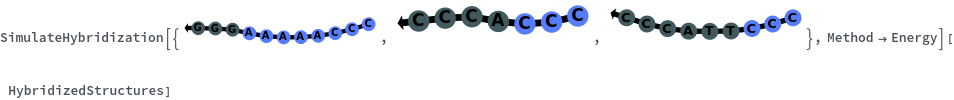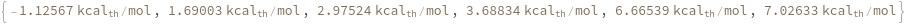SimulateHybridization
SimulateHybridization[oligomer]⟹hybridization
predicts potential secondary structure interactions of the provided list of oligomer.
Details
- This function by default conducts Pairing first with the input pair of oligomers, then conducts Folding to infinity depth to allow intra-molecular pairing. If Folding is turned off in option control, only inter-molecular pairing is considered.
- The function can also handle a list of oligomer lists as input: SimulateHybridization[{{oligomer..}..}].
- Thermodynamic DNA Nearest Neighbor parameters from Object[Report, Literature, "id:kEJ9mqa1Jr7P"]: Allawi, Hatim T., and John SantaLucia. "MeltingCurve and NMR of internal GT mismatches in DNA." Biochemistry 36.34 (1997): 10581-10594.
- Thermodynamic RNA Nearest Neighbor parameters from Object[Report, Literature, "id:M8n3rxYAnNkm"]: Xia, Tianbing, et al. "Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs." Biochemistry 37.42 (1998): 14719-14735.
Input
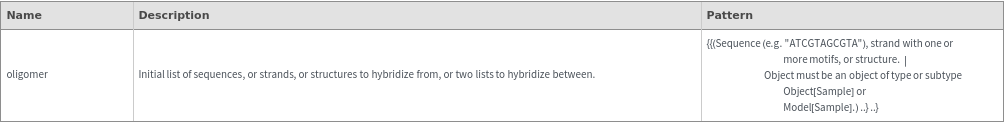
Output

General Options
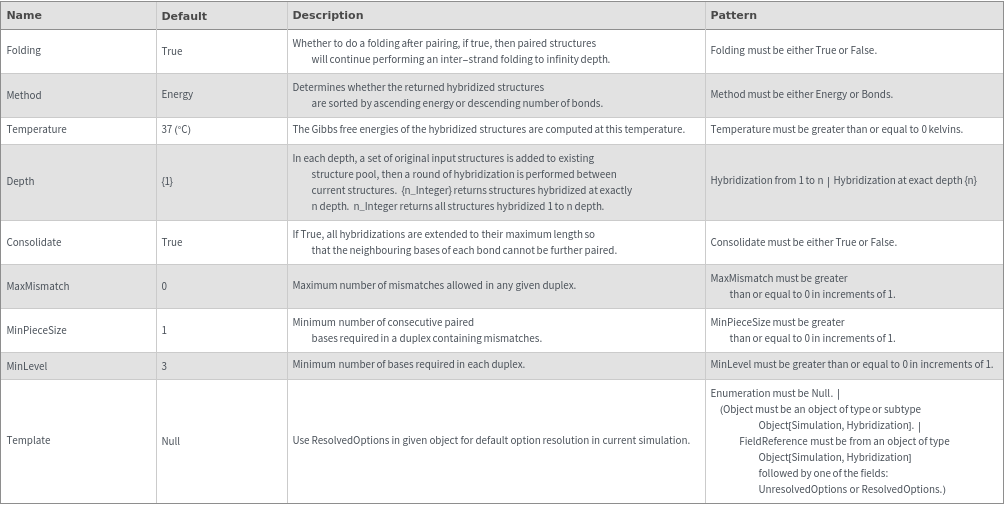
Examples
Basic Examples (5)
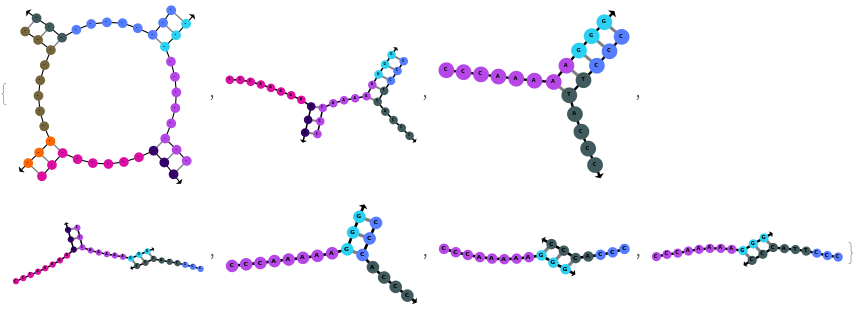
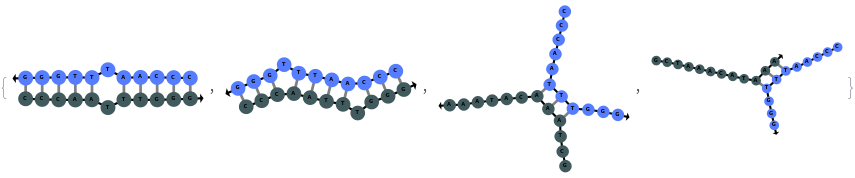
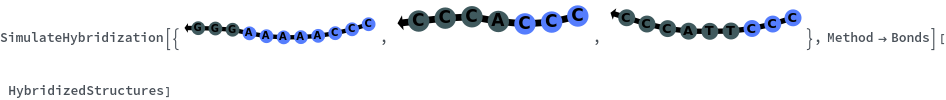
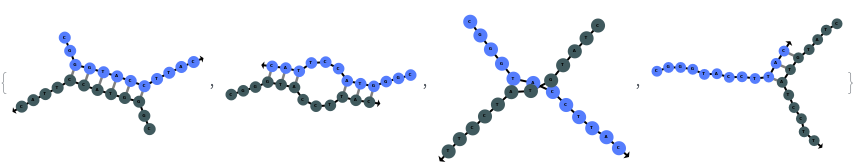
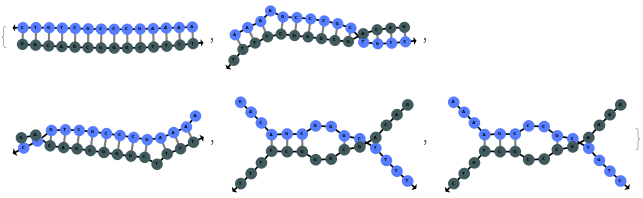
When providing a pair of sequences, SimulateHybridization predicts potentially high energy secondary structures the two sequences can form:

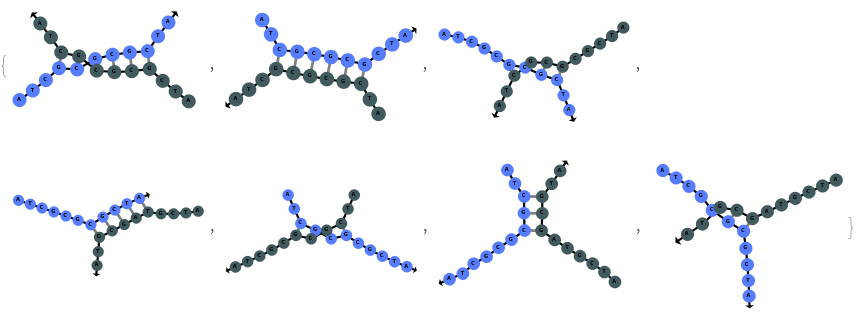
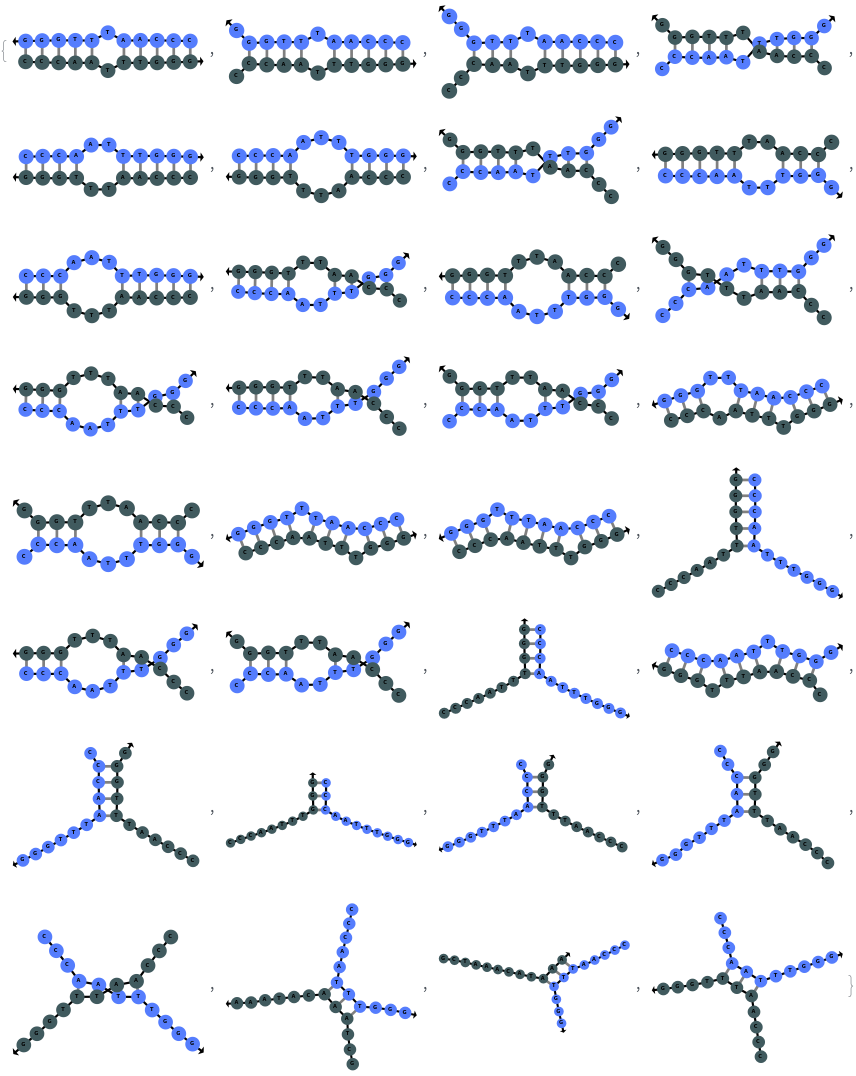
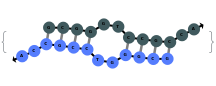
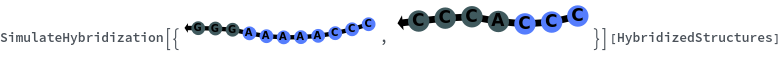
SimulateHybridization can return all possible hybridization results even when the input pair of sequences are perfect Waston-Crick reverse complements:

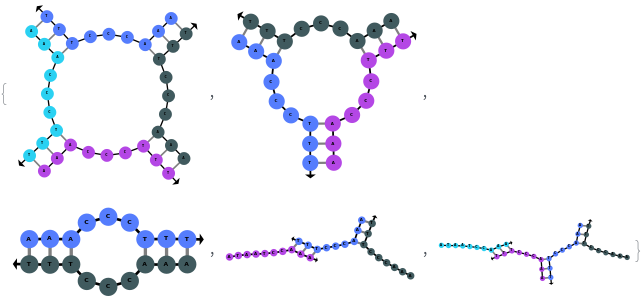
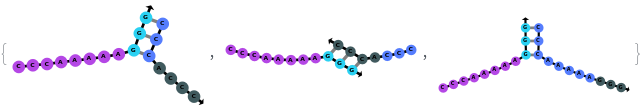
Additionally, SimulateHybridization can handle strands with more than one sequence:
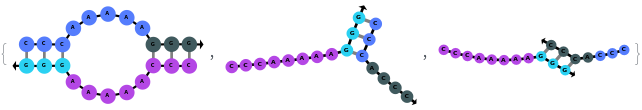
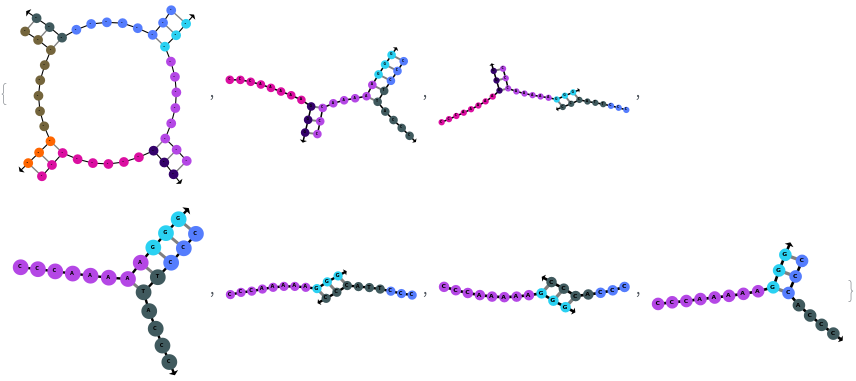
SimulateHybridization can also handle complicate structures:
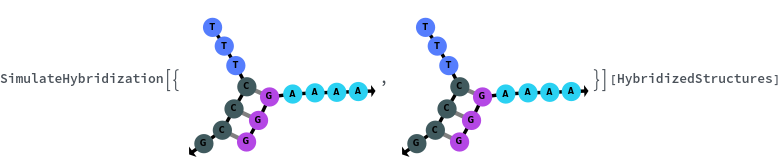
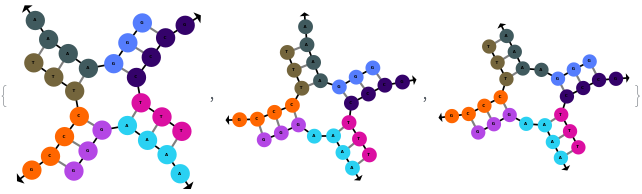
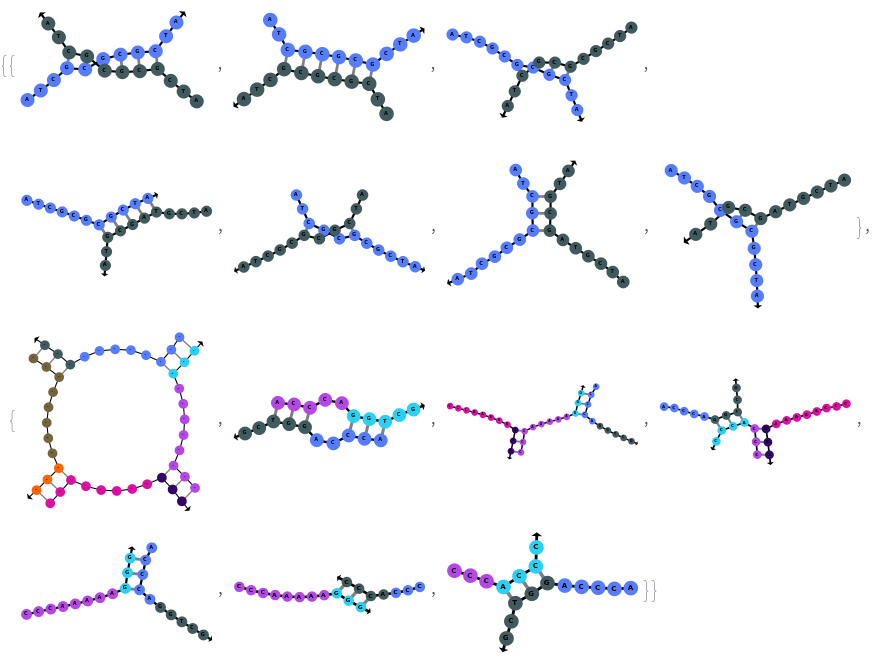
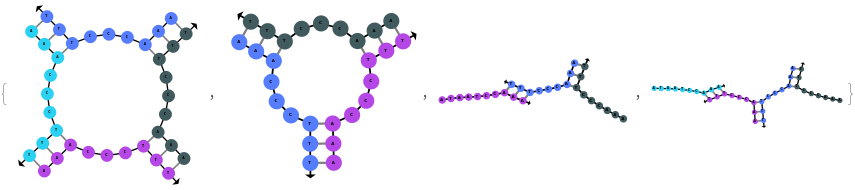
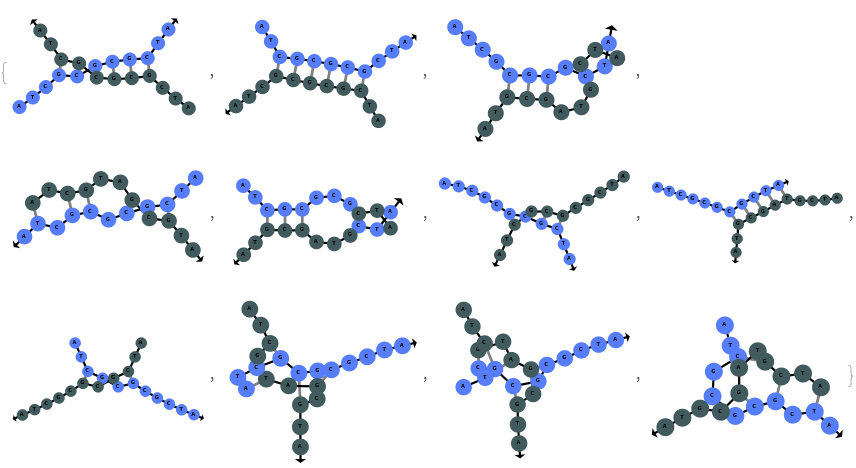
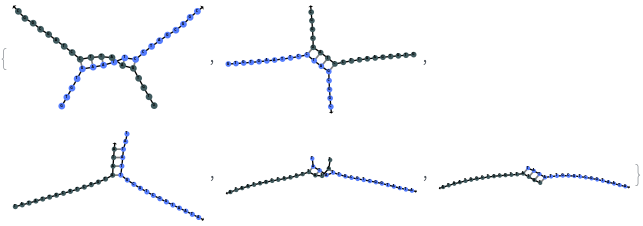
SimulateHybridization can hybridize multiple input oligomers together, returns all possible intermediate and final structures: