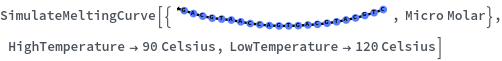
SimulateMeltingCurve
SimulateMeltingCurve[{{initSpecies,initConcentration}..}]⟹thermodynamicCurves
simulates the kinetics of a mechanism generated from a paired set or paired set list of nucleic acids oligomers 'initSpecies' with 'initConcentration' over a range of temperatures specified through options.
SimulateMeltingCurve[primerset]⟹thermodynamicCurves
simulates the kinetics of the molecular beacons that are part of this primerset.
Details
- The function simulates the kinetics of a mechanism generated from an initial set of nucleic acids from HighTemperature down to LowTemperature, then back up to HighTemperature.
- The kinetic mechanism (as stored in the ReactionMechanism field) is generated by SimulateReactivity considering all possible reaction types in NucleicAcidReactionTypeP applied to the initial species given the initial concentrations, then the concentration of each resulting species is calculated as a function of temperature by SimulateKinetics.
- Thermodynamic DNA Nearest Neighbor parameters from Object[Report, Literature, "id:kEJ9mqa1Jr7P"]: Allawi, Hatim T., and John SantaLucia. "MeltingCurve and NMR of internal GT mismatches in DNA." Biochemistry 36.34 (1997): 10581-10594.
- Thermodynamic RNA Nearest Neighbor parameters from Object[Report, Literature, "id:M8n3rxYAnNkm"]: Xia, Tianbing, et al. "Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs." Biochemistry 37.42 (1998): 14719-14735.
Input
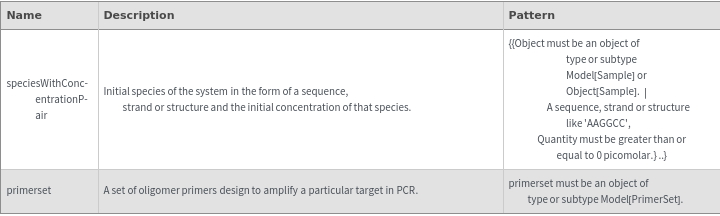

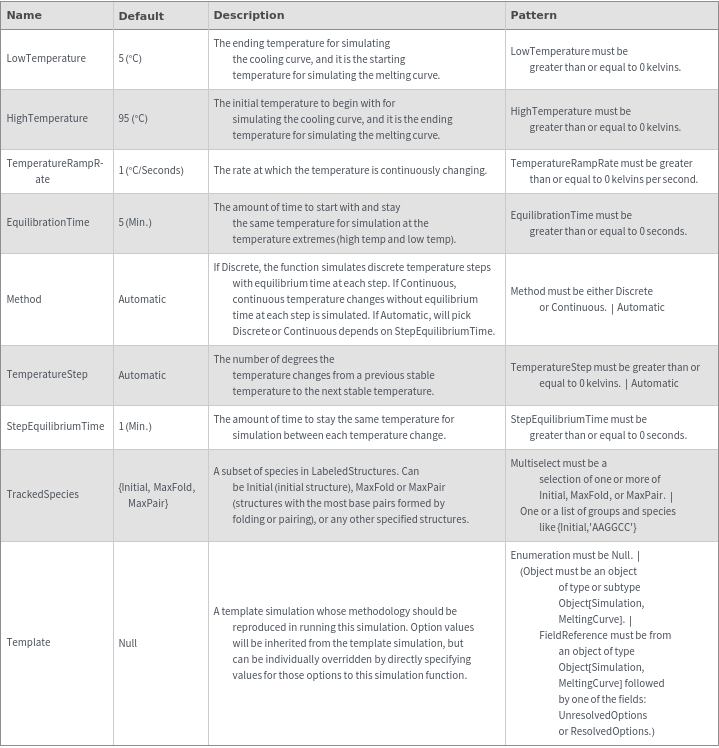
Output
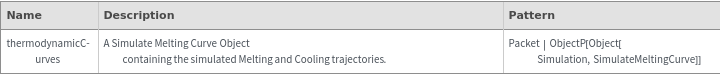
General Options
Examples
Basic Examples (4)
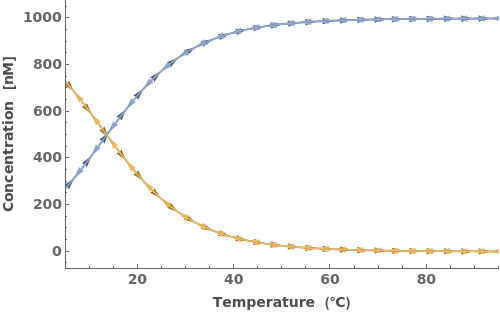
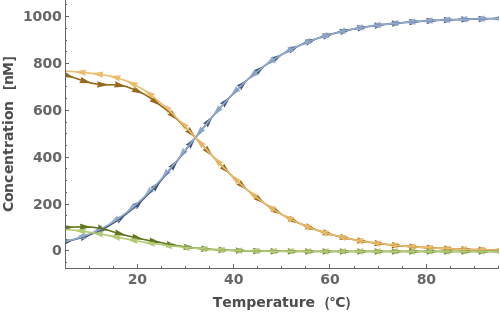
When provided with an oligomer, SimulateThemodynamics simulates the kinetics of a mechanism generated from 'initial' over a range of temperatures (defaults from 5 to 95 celsius):

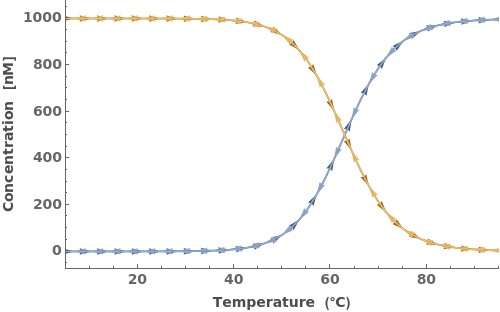
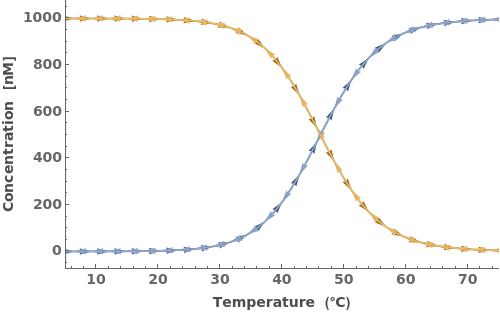
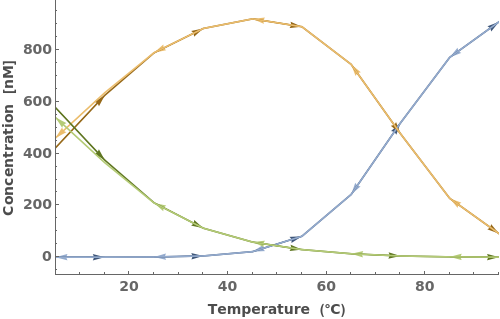
In addition to accepting one oligomer, the function can handle more than one oligomer:

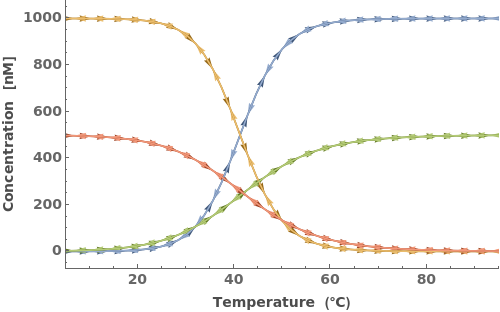
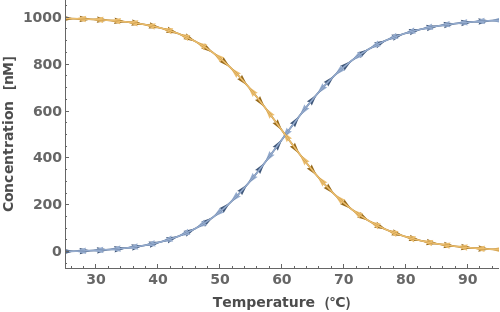
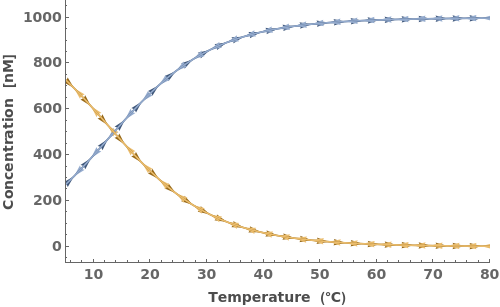
The input oligomers can also be strands or structures:
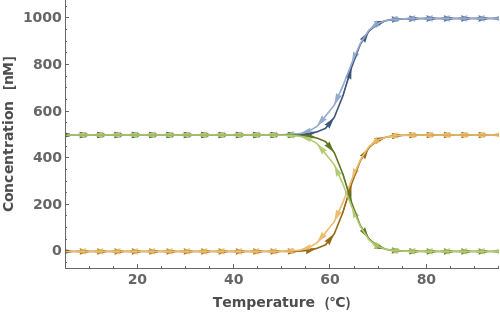
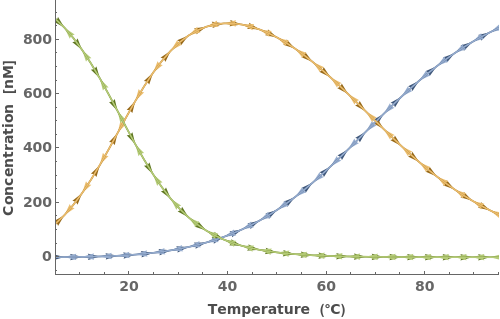
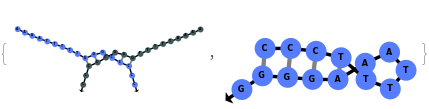
Simulate MeltingCurve for an Object[Sample]:

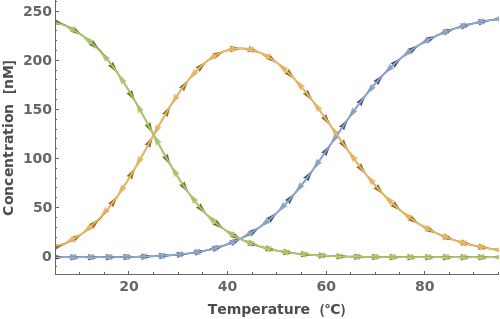
Additional Examples (7)
The function can handle a list of inputs with the form of {sequence, concentration}:

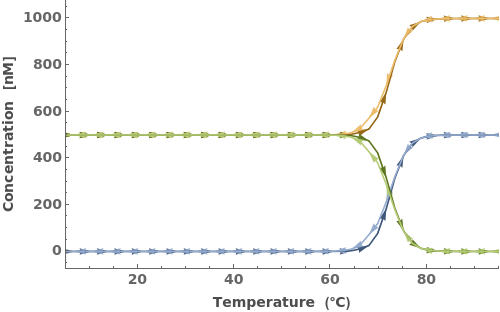
The function can handle a list of inputs with the form of {strand, concentration}:
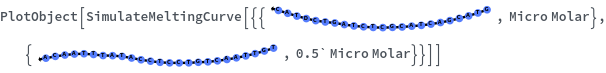

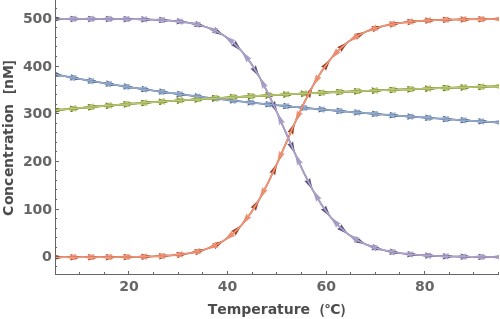
The function can handle a list of inputs with the form of {structure, concentration}:
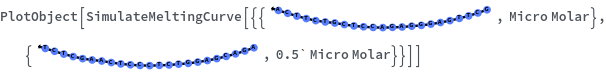

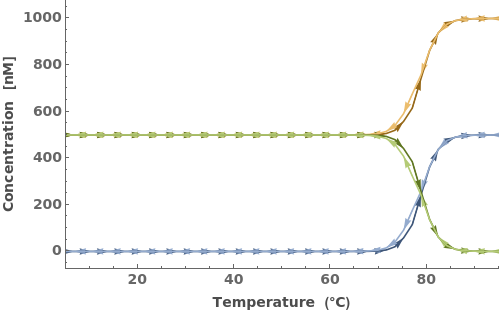
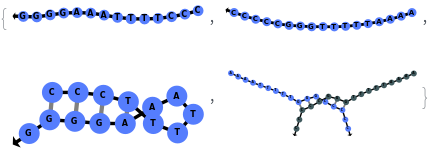
Simulate a beacon folding on itself:

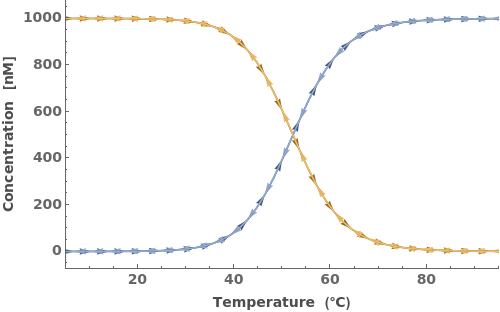
Simulate a beacon binding to its reverse complement:

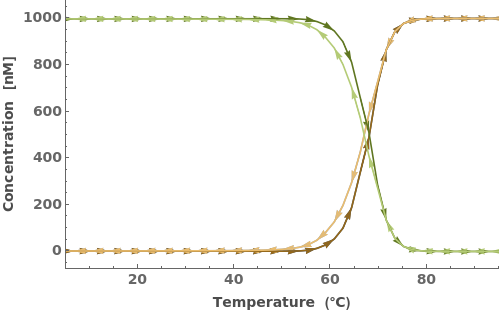
Determine which field(s) to get:

Obtain only MeltingCurve or CoolingCurve if specified:
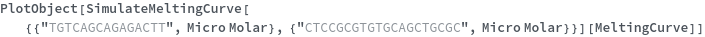
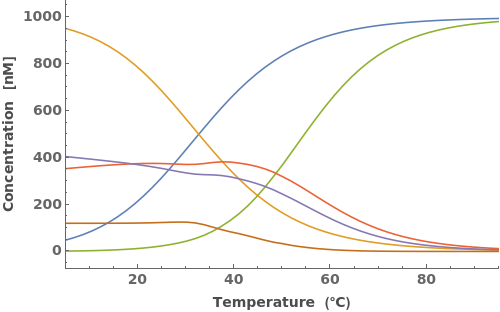
Options (12)
EquilibrationTime (1)
Method (3)
If Discrete, the function simulates discrete temperature steps with equilibrium time at each step:


If Continuous, continuous temperature changes without equilibrium time at each step is simulated:

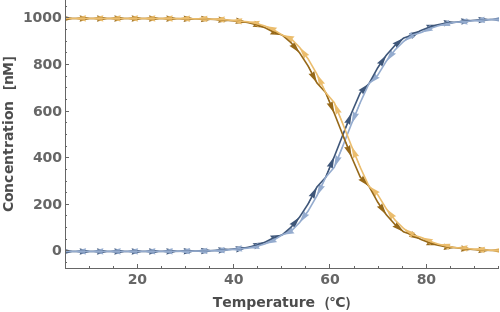
If 'StepEquilibriumTime' is 0, Continuous temperature change is simulated. Otherwise Discrete temperature change is simulated: