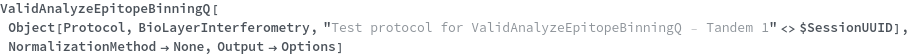ValidAnalyzeEpitopeBinningQ
ValidAnalyzeEpitopeBinningQ[DataObject]⟹TestSummary
returns an EmeraldTestSummary which contains the test results of AnalyzeEpitopeBinning['dataObjects'] for all the gathered tests/warnings or a single Boolean indicating validity.
ValidAnalyzeEpitopeBinningQ[ProtocolObject]⟹TestSummary
returns an EmeraldTestSummary which contains the test results of AnalyzeEpitopeBinning[ProtocolObject] for all the gathered tests/warnings or a single Boolean indicating validity.
Details
Input
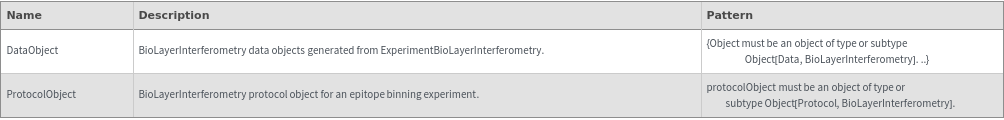
Output

Data Processing Options
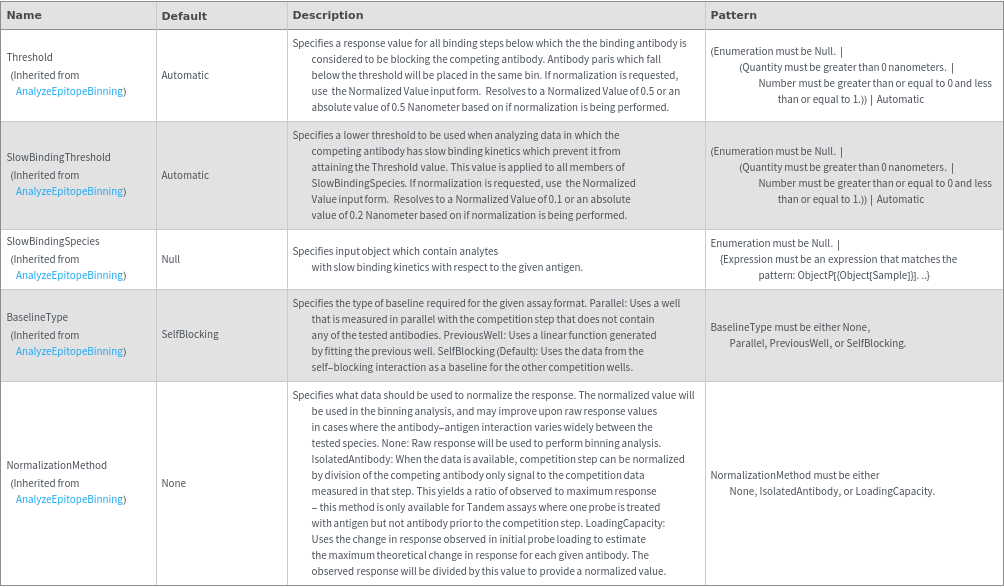
General Options
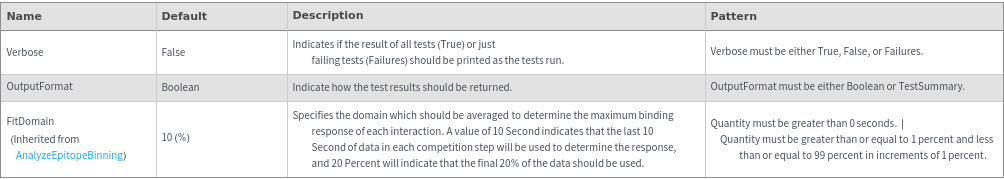

Method Options
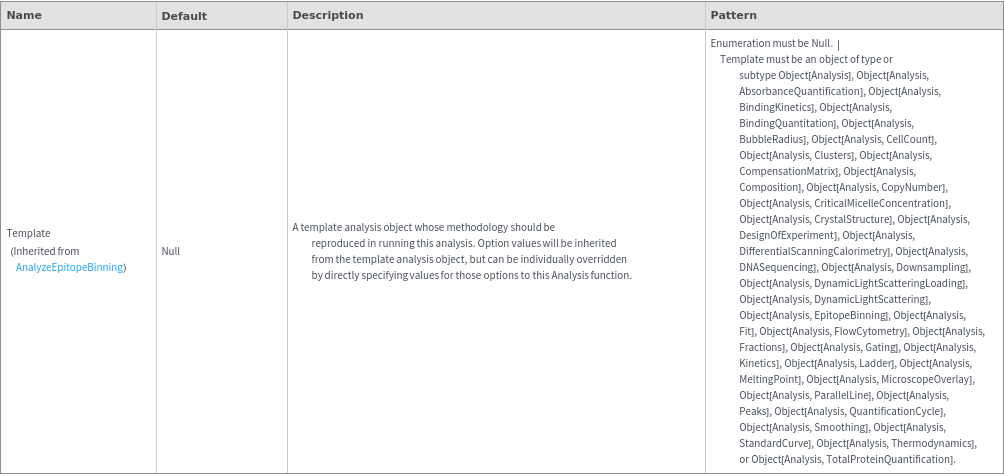
Examples
Basic Examples (2)
Options (11)
BaselineType (2)
Specify the type of baseline well used to account for drift or solvent/media based fluctuation in response signal (PreviousWell) and ValidAnalyzeEpitopeBinningQ returns True:
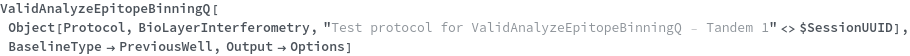

Specify the type of baseline well used to account for drift or solvent/media based fluctuation in response signal (SelfBlocking) and ValidAnalyzeEpitopeBinningQ returns True:


FitDomain (2)
Specify the region of each competition step to average to obtain the response value using percent of the total step time and ValidAnalyzeEpitopeBinningQ returns True:


Specify the region of each competition step to average to obtain the response value using time and ValidAnalyzeEpitopeBinningQ returns True:


NormalizationMethod (2)
Specify the type of well used to normalize response - use a parallel well in which the antibody loads an unblocked antigen covered surface and ValidAnalyzeEpitopeBinningQ returns True:


Specify the type of well used to normalize response - use the initial loading step for a given antibody as the maximum value and ValidAnalyzeEpitopeBinningQ returns True:
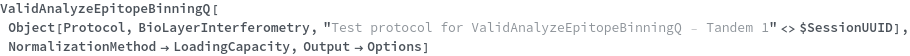

OutputFormat (1)
SlowBindingSpecies (1)
SlowBindingThreshold (1)
Threshold (2)
Automatically resolve the Threshold to a normalized response if the data is analyzed with a NormalizationMethod and ValidAnalyzeEpitopeBinningQ returns True:
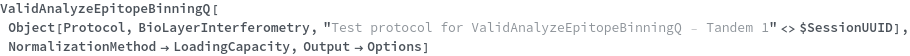

Automatically resolve the Threshold to a raw response (in Nanometer) if the data is not analyzed with a NormalizationMethod and ValidAnalyzeEpitopeBinningQ returns True: