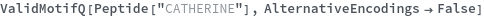ValidMotifQ
ValidMotifQ[motif]⟹out
returns True if motif is a valid motif (explicitly typed sequence).
Examples
Basic Examples (4)
Options (12)
AlternativeEncodings (2)
Degeneracy (4)
If Degeneracy is set to true then degenerate Monomers will be permitted in the motif:


If Degeneracy is set to false, then degenerate Monomers will not be permitted in the motif:


Numeric sequences are rejected when Degeneracy is set to False:


Numeric sequences are accepted when Degeneracy is set to True: