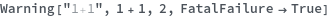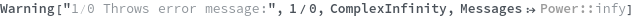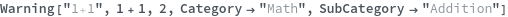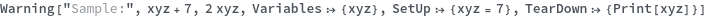Warning
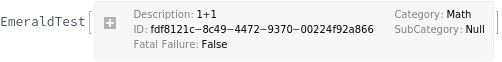
Warning[description,expression,expectedValue]⟹test
returns an executable test which will evaluate expression and determine if it matches expectedValue.
Details
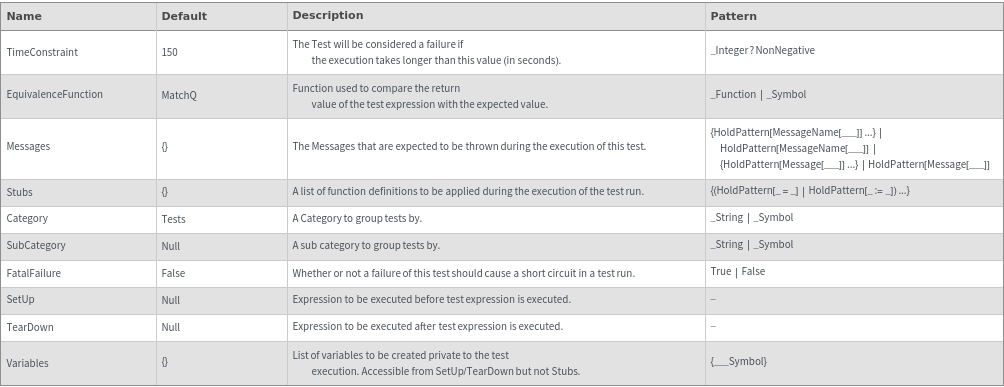
- The following Options MUST be specified as RuleDelayed: Messages,SetUp,TearDown,Variables,Stubs.
- The function definitions specified in the Stubs Options list are only applicable during the execution of the test. During the execution of the test, all previously specified definitions for the stubbed out symbols will be ignored and after the test is executed the stubbed out definitions will be forgotten.
- The 'Stubs' Options differentiates between Set & SetDelayed calls. A SetDelayed call will clear all definitions for the symbol before applying the Stub definition, whereas a Set call will prepend the definition to the existing DownValues for the symbol.
- The 'Set' Stubs (which prepend DownValues) works by copying all the DownValues/OwnValues/UpValues/Options/Attributes for a symbol during a test and as a result will not work with built in Mathematica symbols whose DownValues, OwnValues, etc., are not exposed. However, The SetDelayed method for clearing the symbol will always work.
-
HoldRest -
AbsentTargetMolecules The TargetMolecules `1` are not present in the Data Objects `2`. These Data Objects may need to be analyzed separately or the TargetMolecules should be changed to include a molecule present in this Data. AbsorbanceRateAdjusted The following absorbance sampling rate options, `1`, can only be specific values and were set to the closest one. AbsSpecInsufficientSampleVolume The specified sample volumes `1` are below the minimum required volume for the specified instrument type of `2`. Please consider using larger sample volumes or select an instrument that can read samples of this size. AcquireImageOverlappingDetectionLabels The following objects, `2` ,are specified multiple times as DetectionLabels option at index `1`. No action is needed if this is intentional. AdjustmentDataNotFound No compensation sample was provided for detector(s) `1` and compensation will not be computed for these detectors. If no control samples were prepared for these detectors, this warning may be safely ignored. If control samples were prepared, please ensure that the AdjustmentSamples field of input protocol is correctly linked to all samples, and that the AdjustmentSampleData field contains the corresponding data. AgaroseLoadingDyeRangeCollectionOptionMismatch The following input samples, `1`, have collection-related Options, `2`, whose values, `3`, are not within the range of sizes of Oligomers in the corresponding LoadingDyes, `4`. Most accurate sizing and collection is achieved when the target of interest is between the size of the two LoadingDyes. Please consider specifying loading dyes whose sizes span the desired `2`: AliquotRequired The sample(s) `1`, must be transferred from their current container(s) `2` into `3` `4` AllAbsOutOfRange For sample `1`, all the measured absorbance data are out of the recommended linear range {`2`, `3`}. Continued analyzing while considering them as in-range data. AllWavelengths Since a 3D plot was requested, data for all available wavelengths will be shown. Please set PlotType -> ListLinePlot if you wish to see data at only a specific wavelength. AllWavelengthsOutOfRange All of the specified wavelength values specified are out of the wavelength range that the data was acquired with. Attempting to use 2D curves for this analysis. Please modify the Wavelength option to values in the range from `1` nm to `2` nm. AlphaScreenHighWellVolume The assay volume (AlphaAssayVolume) `1` is close to the maximum working well volume of the plate `2`. AlphaScreenLowWellVolume The assay volume (AlphaAssayVolume) `1` is less than the minimum working well volume of the plate `2`. AlphaScreenNotBMGPlates The plate model `1` doesn't match any available plate layouts in the reader. We recommend to resolve AssayPlateModel automatically or transfer the samples from a prepared plate. AlphaScreenOpticsOptimizationParametersUnneeded When an optics optimization template (OpticsOptimizationTemplate) is used, the options `1` are not needed. The options `1` are ignored and resolved to Null. The Gain and FocalHeight are set based on the template. AlternativeParameterizationNotAvailable The AlternativeParameterization for oligomer `1` does not exist. Setting AlternativeParameterization to False. AmbientTemperatureMeasurement The Samples `1` will be transported at the temperature indicated by the TransportWarmed/TransportChilled fields in their model, but samples will not remain at that temperature during density measurement. AmbiguousAdjustmentSample The provided AdjustmentSample appears multiple times in the input and should be disambiguated to ensure the correct aliquot of the sample is used. By default the first appearance of this sample will be used. Please specify this option in the form {appearance number, sample} if you wish to use another instance. AmbiguousAnalyte The desired analyte for sample(s) `1` is ambiguous because its Analytes or Composition fields contain multiple identity models that could be the desired analyte. Setting to `2`. AmountWillBeMeasured Mass or Volume were not specified for `1`. The amount will be measured upon arrival at the ECL site. This may involve transferring or thawing samples. If you prefer to determine the sample amount yourself, please specify Mass or Volume. AnalytesNotFound The analytes `1` at indices `2` in StandardConcentrations could not be matched to any picked peaks in their corresponding StandardData objects. Missing analyte concentrations in these standards will not be used in the analysis. AnalyzeTPQCalculatedConcentrationsExtrapolated The following members of AssaySamples, `1`, have calculated TotalProteinConcentrations, `2`, that fall outside of the range of concentrations used to create the Standard Curve, between 0 milligrams per milliliter and `3`. The AssaySamples correspond to the InputSamples, `4`, and their calculated TotalProteinConcentrations, `5`. These extrapolated concentrations should be treated with caution, as they are likely to be less accurate than values within the range of the standard curve. If the calculated concentration is larger than `3`, please consider diluting the sample further for future TotalProteinQuantification experiments. AnalyzeTPQConcentrationIncalculable The TotalProteinConcentration of the following AssaySamples, `1`, which correspond to the SamplesIn, `2`, cannot be calculated. Solutions for x (TotalProteinConcentration) to the following equation, y = `3`, when substituting the associated Absorbance or Fluorescence values as y, `4`, do not exist or are negative. This can occur for Sigmoidal standard curve fitting when the TotalProteinConcentration of the AssaySample is outside of the dynamic range of the assay, or for some Cubic fits. To obtain concentration information, please set the StandardCurveFitType to Linear. Alternatively, the TotalProteinConcentration can be measured by running ExperimentTotalProteinQuantification on input samples that are more dilute. AnalyzeTPQPoorStandardCurveFit The fit analysis, `1`, has an R-squared value, `2`, which is lower than 0.95. This analysis was performed at the following QuantificationWavelength(s), `3`, using `4` as the StandardCurveFitType. It is possible that the QuantificationWavelength used was not ideal. Please consider trying other QuantificationWavelengths, or changing the StandardCurveFitType. APIConnection A connection to the scraping API was not able to be formed. Please try re-running this function again and check your firewall settings. AreaToleranceExceeded SplittingRelativeIntegralTolerance exceeded while assigning multiplicity `1` to peaks at `2` (relative areas: `3`). Please verify that the multiplicity assignment is correct, or increase SplittingRelativeIntegralTolerance. ArrayCardExists An array card with these primers and probes already exists. Please use `1` directly in ExperimentqPCR. AutoclaveContainerTooFull The following containers, `1`, have Contents with Volumes larger than 3/4 of the MaxVolume of the container. We will attempt to move the contents into the following containers, `2`, so that they can be autoclaved. If this is not desired, please choose a different input container or transfer the contents to a desired container. AutomaticallySelectedDilutionSolvent No `1`DilutionSolvent has been specified, using default Hexanes. AutomaticallySelectedWashSolvent No `1`SyringeWashSolvent has been specified, using default Hexanes. AutomaticAssignmentNotAvailable Automatic assignment of NMR peaks to molecular structure is not available for nucleus type `1`. Peaks must be annotated manually using option SplittingAssignments. AutomaticSampleGiven The Sample option wasn't specified at the current LabelSample primitive. The LabelSample primitive requires a Sample to be specified (either a new Model[Sample] to transfer into the container or an existing Object[Sample] already in the container). If you wish to label an empty container, use the LabelContainer primtiive instead. The Sample option will default to Model[Sample, "Milli-Q water"]. autoNeutralLossValueWarnings For samples `1` in ESI-QQQ using NeutralIonLoss as the ScanMode, a valid mass value in NeutralLoss option needs to be specified in order to generate useful and reliable data. The experiment can further procees with an automatically assigned value (50 gram/mole), but this experiment may not generate any usefule results. autoResolvedFragmentMassDetectionFixedValues For samples `1` in ESI-QQQ using `2` as the ScanMode, a single mass value (for PresursorIonSacn mode only) or a list of mass values (for MultipleReactionMonitoring mode) should be specified in FragmentMassDetection option. The experiment will proceed with FragmentMassDetection being resolved based on MassDetection or was arbitrarily set to 500 gram/mole. Proper mass values for this option needs to be specified to collect useful data. autoResolvedMassDetectionFixedValue For samples `1` in ESI-QQQ using `2` as the ScanMode, a single mass value (for ProductIonSacn mode only) or a list of mass values (for MultipleReactionMonitoring and SelectedIonMonitoring mode) should be specified in MassDetection option. The experiment will proceed with MassDetection being resolved based on samples' molecular weights (if exist) or was arbitrarily set to 500 gram/mole. Proper mass values for this option needs to be specified to collect useful data. BadObservableSpecies Selected observable species `1` is invalid, so defaulting to `2` as observable species. BadPolymerType Given polymer type `1` does not match on input in function `2`. Polymer will be defaulted to Automatic. BedVolumeOverride As the BedVolume has been specified the, ResinLoadingAmount will be overridden from the backcalculation of the BedVolume and the value of the SpecificVolume for the set resin and storage buffers, `1`. BillNotFinished Input Bill has Status of Open, please keep it in mind. BindingKineticsAnalysisMissing There is no BindingKinetics analysis object linked to the input data object(s). If you would like the data to be fit, please run AnalyzeBindingKinetics on the data objects. BindingQuantitationAnalysisMissing There is no BindingQuantitation analysis object linked to the input data object(s). If you would like the data to be fit, please run AnalyzeBindingQuantitation on the data objects. BindingQuantitationConflictingDomainAndParameter The domain specified by `1` is inconsistent with the selected FittedParameter. When fitting InitialRate, use InitialFitDomain, and when fitting AverageEquilibriumResponse, use FinalFitDomain. BlankStateWarning The blanks (`1`) do not have a state of liquid. If this is not intended, please check the blank samples (`1`). BLIActivateWithoutLoad The Activate step is generally selected along with a Load step. Select Load in LoadingType if you wish to load the probe surface, and populate the other required parameters. BLIConflictingStartDelayOptions The options `1` are in conflict. If a time is specified for the start delay, also specify if the plate should be shaken during this time. BLIInstrumentPrecision The value `2` in the field `1` has been rounded to match instrument precision. BLIInsufficientDevelopmentFixedDilutionsVolume The DetectionLimitFixedDilutions require transfer volumes below the achievable volume of 1 Microliter for the following samples: `1`. BLIInsufficientDevelopmentSerialDilutionsVolume The DetectionLimitSerialDilutions require transfer volumes below the achievable volume of 1 Microliter for the following samples: `1`. BLIInsufficientKineticsFixedDilutionsVolume The KineticsSampleFixedDilutions are less than 210 Microliter for the following samples: `1`. The recommended range is 210 Microliter to 250 Microliter. BLIInsufficientKineticsSerialDilutionsVolume The KineticsSampleSerialDilutions are less than 210 Microliter for the following samples: `1`. The recommended range is 210 Microliter to 250 Microliter. BLIInsufficientPreMixVolume The PreMixSolutions are less than 250 Microliter for the following samples: `1`. The recommended range is 250 Microliter to 220 Microliter. BLIPlateCoverNotRecommended The total predicted assay time of `1` does not exceed 4 hours. A plate cover is generally not required in this case. BLIPlateCoverRecommended The total predicted assay time of `1` exceeds 4 hours. It is recommended that PlateCover be set to True to avoid excessive evaporation, leading to changes in solution concentration during the assay. BLIPrimitiveOverride User input AssaySequencePrimitives or ExpandedAssaySequencePrimitives will override other options, except ExperimentType and those related to dilutions such as KineticsSampleSerialDilutions. BLIPrimitiveValueInstrumentPrecision The value `1` for field `2` in the `3` primitive has been rounded to match the instrument precision. BLIProbeApplicationMismatch The BioProbeType (`1`) is intended for ExperimentType -> `2`, and may not be suitable for `3`. Validate your assay to confirm that the probe functions properly for this application if you are unsure. BLIProbeRegeneration The BioProbeType (`1`) may not be suitable for regeneration for ExperimentType -> `2`. Validate that the probe can be regenerated for `2` if you are unsure. BLIQuenchWithoutLoad The Quench step is generally selected along with a Load step. Select Load in LoadingType if you wish to load the probe surface, and populate the other required parameters. BLIRegenerationNotSpecified For the option RegenerationType, PreCondition must be set along with any or all of the following: Neutralize, Wash, Regenerate. BLIUnusedAmplifiedDetectionOptions The following options are only used if AmplifiedDetection is selected in QuantitationParameters: `1`. BLIUnusedBlank Blank was specified but not used. Specify change the value of `1` for the Blank solution to be measured in parallel with the samples. BLIUnusedQuantitationEnzymeOptions The following options are only used if EnzymeLinked is selected in QuantitationParameters: `1`. BLIUnusedQuantitationStandardOptions The following options are only used if StandardWell or StandardCurve is selected in QuantitationParameters: `1`. BLIUnusedQuantitationStandardWellOptions The following options are only used if StandardWell is selected in QuantitationParameters: `1`. BLIUnusedStandard Standard was specified but not used. Specify Standard in `1` for the Standard solution to be measured in parallel with the samples. BothSDSBufferOptionsSet Both Concentrated SDS buffer and SDS buffer options were specified for `1`. Please consider setting one or the other. By default, the Concentrated SDS Buffer option will be used when both options are specified. BottomReadingAliquotsRecommended For the best results, it is recommended to aliquot the samples into a new plate with WellColor of Clear when ReadLocation is set to Bottom. BufferConflict The resolved values of `1` (`2`) differ from provided gradient method(s) for these options (`3`). Nonetheless, the experiment will commence as directed with the former value(s). BufferWillNotBeUsed AssayBuffer and/or ConcentratedBuffer were specified for `1`. However, the buffer(s) will not be used because AssayVolume and Volume are equal and no dilution will occur for these samples. CalculatedBedVolume As the ResinLoadingAmount has been specified and the BedVolume has not been, the BedVolume will be backcalculated from the ResinLoadingAmount and the value of the SpecificVolume for the set resin and storage buffers, `1`. CalculatedResinLoadingAmount As the BedVolume has been specified, the ResinLoadingAmount will be backcalculated from the BedVolume and the value of the SpecificVolume for the set resin and storage buffers, `1`. CalibrantMassDetectionMismatch For the following input samples, `1`, the mass range that is chosen or automatically determined, `2`, does not fully lie inside the calibrant's `3` mass range, which may result in suboptimal calibration for the masses outside the calibrant's mass range. Consider adjusting the mass range or choosing a different calibrant. Please refer to the documentation for a list of suitable combinations of mass range and calibrants. CalibrantSurfaceTensionMismatch The SurfaceTension field of the calibrant `1` does not match the value of the specified CalibrantSurfaceTension `2`. CannotCollapse `1` has not been collapsed because it does not consist entirely of the same value. This option has been left in index-matched form. CannotImageFluorescentDetectionLabels The instrument is not capable detecting fluorescent DetectionLabels contained by the following input samples: `1`. Please specify a new instrument that supports ConfocalFluorescence or Epifluorescence mode if detection of fluorescent DetectionLabels is desired. CannotImageNonFluorescentDetectionLabels The instrument is not capable detecting non-fluorescent DetectionLabels contained by the following input samples: `1`. Please specify a new instrument that supports BrightField or PhaseContrast mode if detection of non-fluorescent DetectionLabels is desired. CannotTransform3DData The x-axis of the data cannot be transformed if the data is 3D. Plot will be generated without the transform. CannotTransformMassSpecData Cannot transform the x-axis of the mass spec data to be volume. Plot will be generated without the transform. CapillaryELISACartridgeExist The requested Capillary ELISA cartridge already exists. Please use `1` directly in ExperimentCapillaryELISA. CapillaryELISASampleDilution Dilution is required on all the loading samples for the best performance of microfludics. While `1` is set to Null for the samples `3`, dilution is automatically selected using `2`. To directly use the sample without dilution, please consider setting DilutionCurve to FixedDilutionVolume with DiluentVolume of 0 Microliter. CapillaryELISAStandardDilution Dilution is required on all the loading samples for the best performance of microfludics. While `1` is set to Null for the standard samples `3`, dilution is automatically selected using `2`. To directly use the standard sample without dilution, please consider setting StandardDilutionCurve to FixedDilutionVolume with DiluentVolume of 0 Microliter. CaptureAntibodyDilutionRecommended CaptureAntibodyDilution is recommended (not Null) for capture antibody samples `1` (used for ELISA assay for samples `2`), because resupension and/or conjugation of the antibody is performed in this protocol and the concentration of the antibody may be too high for capillary ELISA experiment. CellCountNoData There is no data provided in the following protocol(s): `1`. CellCultureTypeNotSpecified The sample(s), `1`, have no CellCultureType specified in the options or Object. For these sample(s), the CellCultureType is defaulting to Suspension. If this is not desired, please specify a CellCultureType CellTypeNotSpecified The sample(s), `1`, have no CellType specified in the options or Object. For these sample(s), the CellType is defaulting to Microbial. If this is not desired, please specify a CellType CentrifugePrecision The specified intensities `1` are not attainable by the precisions `2` of the centrifuge that will be used. Therefore, these intensities have been rounded to `3`. CentrifugeRecommended Because the following samples will have their volumes Ultrasonically measured, we recommend preparing the samples with Centrifuge -> True to reduce errant volume measurements caused by material left on the sides of the container: `1`. CIEFMakeMasterMixAnodicSpacerComposition The composition of the AnodicSpacer object/model does not have a known anodic spacer (e.g., Iminodiacetic acid) for `1`. Volumes will be calculated assuming the anodic spacer's stock solution concentration is 200 mM. Consider specifying a different object/model or specify the volume to be added to the master mix. CIEFMakeMasterMixCathodicSpacerComposition The composition of the CathodicSpacer object/model does not have a known cathodic spacer (e.g., arginine) for `1`. Volumes will be calculated assuming the cathodic spacer's stock solution concentration is 500 mM. Consider specifying a different object/model or specify the volume to be added to the master mix. CIEFMakeMasterMixDenaturationReagentComposition The composition of the DenaturationReagent object/model does not have a known denaturation agent (e.g., SimpleSol and Urea) for `1`.Volumes will be calculated assuming the stock solution concentration is 10 M. Consider specifying a different object/model or specify the volume to be added to the master mix. CIEFMakeMasterMixElectroosmoticFlowBlockerComposition The composition of the ElectroosmoticFlowBlocker object/model does not have a known elerctroosmotic flow blocking agent (e.g., methyl cellulose) for `1`. Volumes were calculated assuming a 1% concentration. CIEFMissingSampleComposition Sample composition is missing for `1`. As a result, sample Volume can't be accurately calculated to reach 0.2 mg/mL protein and defaulted to 10% of the TotalVolume, or to 25 microliters if OnBoardMixing is True. Please specify the volume if desired. CIEFPreMadeMasterMixWithMakeOwnOptions Options for both premade master mix and modular master mix are specified for `1`. These options are exclusive to eachother and only premade master mix options will take effect. If you don't wish to use a PremadeMasterMix, make sure to set this option to False. CIEFresolveableDenatureReagentConcentrationUnitMismatch The DenaturationReagent target concentration unit (VolumePercent) does not match that of the DenaturationReagent object/model for `1`. The volume has been calculated considering the reagent's concentration as 100%. Please consider specifying the target concentration in Molar concentration. CircularDichroismEnantiomericExcessWavelengthsNotCovered For samples `1`, its detection wavelengths `2` cannot fully cover its specified wavelengths for EnantiomericExcessWavelengths (`3`). Please use wavenlengths that will be detected during the experiement (in the range of DetectionWavelength). CircularDichroismInconsistentAnalyteConcentration For samples `1`, the specified AnalyteConcentration `2` are not same as the concentrations `3` that are stored in samples' composition. Please double-check to make sure correct concentrations are specified. CircularDirchroismUnknownAnalytes For samples `1`, the analyte `2` are not present in sample models composition. Please make sure the specified Ananlyte is desired. CoatingButNoBlocking Coating is True but Blocking is False. We recommend setting Blocking to True for the best ELISA result. ColumnLoadingsDiffer The amount of active sites `1` available on `2` are not equal. Be advised that options that automatically resolve based on scale may not be optimal for the amount of resin used. The experiment will proceed with the specified options. CompanyRequiredForModelInputs These model sample inputs `1` must be provided alongside a service company input. Please specify the service company. Alternatively, specify the samples as a product if you do not want to specify the service company. CompensationMatrixNotFound The parent protocol `1` of object `2` has CompensationSamplesIncluded set to True, but no CompensationMatrixAnalyses. No compensation will be used in this analysis - if compensation is desired, please run AnalyzeCompensationMatrix on parent protocol and then re-run this analysis. ComponentOrder The provided formula adds an acid before the rest of the liquid components. Without a manual override, the component addition order will be changed to `1` such that the acids are added after other liquid components. If you are certain that adding the components in the order you have specified will not present a safety hazard, you may certify that this is a true statement using the SafetyOverride->True argument, which will then revert bask to the order of additions as you have specified it. ComponentStateUnknown The formula component `1`'s State is unknown; the component's amount has not been validated. ComputationNotAborted The computation(s) `1` have status `2` and could not be aborted. Computations must have a status of Queued, Ready, Staged, or Running to be Stopped. ComputationNotFound The computation(s) `1` could not be found in Constellation. Please ensure these objects exist, and that you have permissions to access them. ComputationNotStopped The computation(s) `1` have status `2` and could not be stopped. Computations must have a status of Queued, Ready, Staged, or Running to be Stopped. ConflictAntibodyEpitopes The CustomCaptureAntibody and CustomDetectionAntibody of the samples `1` share the same binding epitopes on the analytes. The ELISA results may be affected. Please consider using different antibody pairs. ConflictBoxWhisker The BoxWhiskerType has been set to Null for Histogram plot. If a BoxWhiskerChart is required, please set PlotType to BoxWhiskerChart. ConflictCapillaryELISAStandardComposition StandardComposition concentration of standard samples `1` should be the same as StandardResuspensionConcentration when resuspension of solid sample is performed in this protocol. ConflictFractionOptionSpecification The specified options `1` conflict with the resolution of no fraction collection. These options will be ignored. ConflictingAlgorithms Image processing algorithms `1` and `2` for input `3` can not be True at the same time. The latter one will be set to False. Please use either one of the algorithms or explicitly provide ImageAdjustment and ImageSegmentation options. ConflictingCalibrantion The specified CalibrationStandard and CalibrationConductivity conflict.Please change the value of these options or let them be automatically resolved. ConflictingCellRadiusOptions Specified MinCellRadius `1` is greater than specified MaxCellRadius `2`. The issue might be due to using one of the MinCellRadius or MaxCellRadius with the other one set to their default value. In this case, both values will be set the same value. ConflictingCleaningSolution A CleaningSolution is given `2` but no CleaningMethod that uses a Solution is specified. ConflictingComponentRadiusOptions Specified MinComponentRadius `3` is greater than specified MaxComponentRadius `4` for input `1` image `2`. It will be set to the default value. ConflictingDefaultCellRadius Either MinCellRadius or MaxCellRadius for input `1` has been specified which is incosistent with the default cell radius settings. MinCellRadius `2` needs to be less than MaxCellRadius `3` to perform PropagateUncertainty, therefore, MinCellRadius and MaxCellRadius is not utilized. Please either adjust the value of the currently specified option or specify both MinCellRadius and MaxCellRadius explicitly. ConflictingDiluents The input data objects have conflicting diluents `1`. Please choose data objects with the same diluent for more accurate analysis. ConflictingDilutionMixVolume DilutionMixVolume `1` is specified but no there is no dilution in the DilutionCurve `2` for the samples `3`. ConflictingOptionsWithinContainer Option values specified for sample(s) `1` are conflicting with other samples in the same container. These samples will therefore be transferred into a new container. ConflictingPeakAssignments The PeakAssignments to the maximal-area peak in ranges `1` result in assignment to the same peak. The last PeakAssignment for these ranges will be used. Please modify the peak ranges so that only one assignment is made to each peak. ConflictingPreparedPlateCalibrants A PreparedPlate is specified `1` and the samples in the calibrant wells `2` do not have the same composition. ConflictingVerification The specified VerificationStandard and VerificationConductivity conflict.Please change the value of these options or let them be automatically resolved. ConflictingVolumeCheckSamplePrepOption The VolumeCheckSamplePrep option was specified True, but PreparatoryPrimitives were not specified. VolumeCheckSamplePrep option will be ignored. ConflictStandardAntibodyEpitopes The StandardCaptureAntibody and StandardDetectionAntibody of the standard samples `1` share the same binding epitopes on the analytes. The ELISA results may be affected. Please consider using different antibody pairs. ConsequtiveSameStorage The consolidated StorageSchedules of objects `1`, contains consecutive entries pointing to the same storage condition. This is typically not necessary. ContainerCentrifugeIncompatible The samples `1` are in containers that cannot fit on the centrifuges `2`. These samples will be transferred to `3`. ContainerOutNotValidated The volume of `1` is not known. If the volume of the sample exceeds the volume of the specified container out, `2`, the sample will be transferred to a different container. ContainersIncludeAdditionalSamples Your input samples occupy a container that contains other samples that are not designated for shipping: `1`. The samples will be transferred to separate containers. ContainersSpanShipments In order to ship samples with the provided ShippingSpeed and ColdPacking specifications, samples that currently occupy the following containers, will be transferred to separate containers. ContainerWithNonInputSamples The following containers also contain samples other than the input samples for the experiment: `1`. These samples may be affected while performing manipulation in an acoustic liquid handler. CorrectionCurveNotMonotonic The specified CorrectionCurve has actual values that are not monotonically increasing. CosolventConflict The resolution of `1` differ from provided gradient methods' respective cosolvents. Nonetheless, the experiment will commence as directed. CoveredTopRead When RetainCover->True bottom reads are recommended to prevent the plate cover from interfering with the readings. Please consider using ReadLocation->Bottom. CriticalMicelleConcentrationAnalysisMissing There is no AnalyzeCriticalMicelleConcentration analysis object linked to the input SurfaceTension data object(s). If you would like the data to be fit, please run AnalyzeCriticalMicelleConcentration on the data objects. CrossFlowAssumedDensity Since the`1` density of the solvent`2` is unknown, it was assumed to be `3` based on a volumetrically weighted average of the liquid components. If this density is inaccurate, please specify the density of the sample, or specify Targets in volume or fold sample volume units. CrossFlowDefaultFilterReturned Since a filter with required sterility, volume range and size cutoff could not be found, a default filter will be returned. If this filter is not acceptable for this experiment, please specify a different SizeCutoff or a CrossFlowFilter. CrossFlowDiafiltrationBufferAssumed Since no DiafiltrationBuffers were specified, water was assumed as the DiafiltrationBuffer. Please specify DiafiltrationBuffers if water is not the desired buffer. CrossFlowDiafiltrationBufferIgnored A diafiltration buffer was specified for a concentration step. The specified buffer `1` will be ignored. CrossFlowDiafiltrationTubingIgnored TubingType was specified for the diafiltration buffer without a diafiltration step. The specified TubingType for the diafiltration buffer `1` will be ignored. CrossFlowFilterSizeCutoffUnavailable A filter with required sterility, volume range and specified SizeCutoff `1` could not be found. A filter with the SizeCutoff `2` will be returned. If SizeCutoff `2` is not acceptable for this experiment, please specify a different SizeCutoff or a CrossFlowFilter. CrossFlowFilterSpecifiedMismatch The specified filter `1` does not match specified option `2`, whose value is `3`. The specified option `2` will be ignored. CrossFlowFilterSterileUnavailable Even though the experiment resolved to Sterile, a sterile filter with the specified SizeCutoff `1` could not be found. A non-sterile filter with the specified cutoff will be returned. If a non-sterile filter is not acceptable for this experiment, please specify a CrossFlowFilter. CrossFlowInstrumentalPrecision The specified precision for option(s) `1` is beyond the capabilities of the instrument. The values of these options will be rounded to instrumental specifications of `2`. CrossFlowLongScanTime The specified absorbance wavelength scan cannot be completed within AbsorbanceDataAcquisitionFrequency. The scan will be completed as requested, albeit less frequently than AbsorbanceDataAcquisitionFrequency. CrossFlowMissingVolumeInformation Sample "`1`" is missing volume information. The maximum volume of the container will be assumed as the volume of the sample. CrossFlowNonSterileSampleReservoir Resolved sample reservoir `1` is not available in a sterile form. If sterility is necessary beyond standard packaging for this experiment, please specify a sterile sample reservoir. CrossFlowNoSampleSpecified No sample is specified. No protocol objects will be generated. CrossFlowRetentateVolumeCapped The retentate volume generated in this experiment is above the capacity of the specified RetentateContainerOut (`1`). The RetentateOutVolume has been capped to the maximum of the specified RetentateContainerOut (`1`). This means some of the retentate will be discarded. If you would like to keep all of the retentate, specify a larger RetentateContainerOut. CrossFlowSampleVolumeExceedsCapacity The specified sample `1` has a volume that exceeds the capacity of the experiment (`2`). The sample volume will be adjusted to `2`. Data2D Since the data is two dimensional, PlotType will be set to ListLinePlot. DefaultFitTypeUndefined No default fit type is associated with standard data type(s) `1`. FitType has been defaulted to Linear. DefaultMassDetection For samples `1`, the molecular weight is unknown and the sample type is generic. As a result a default mass range of 350-2000 m/z will be chosen, which may not cover the mass of the analyte(s) of interest. If you have a sense of the expected mass(es), please consider providing a custom mass range or informing the molecular weight of the samples. DegasSpargingTimeLow In ExperimentDegas, the resolved SpargingTime (`1`) is too low for samples (`2`) and may result in insufficient degassing during sparging. DegasVacuumTimeLow In ExperimentDegas, the resolved VacuumTime (`1`) is too low for samples (`2`) and may result in insufficient degassing during vacuum degassing. DegenerateMassUncertain Exact mass is not defined for degenerate monomers `1` of type `2`. Isotopic variation will be ignored for these monomers and their average molecular weight will be used instead. DeprecatedModel The object `1` is deprecated and results may be incorrect. Please specify a physics model which is still in use. DeprecatedProduct The following model(s) that were requested have no non-deprecated products associated with them: `1`. In the event that all is consumed before the protocol is run, ECL will not be able to order more and may abort this protocol. DeprecatedSpecifiedModels The following model object(s) specified are deprecated: `1`. Please check the Deprecated field for the models in question, or provide non-deprecated models. DeprecatedTemplate The provided Template option `1` is deprecated; it will not be used for default option resolution. DetectionAntibodyDilutionRecommended DetectionAntibodyDilution is recommended (not Null) for detection antibody samples `1` (used for ELISA assay for samples `2`), because resupension and/or conjugation of the antibody is performed in this protocol and the concentration of the antibody may be too high for capillary ELISA experiment. DetectionLabelsNotSpecified The DetectionLabel option at index `1` is not specified while the given Mode will perform fluorescence imaging. All related options will be automatically set to default values if not specified. No action is needed if this is intentional. DeveloperOnlyOptions Current user does not have adequate permissions to set option(s) `1`. These option(s) will be ignored and reverted to their defaults. DFAAgitationSamplingFrequencyLow In ExperimentDynamicFoamAnalysis, the AgitationSamplingFrequency selected (`1`) is low, and would cause data to be sampled only once during the specified AgitationTime (`2`) for this experiment: `3`. DFAAgitationTime In ExperimentDynamicFoamAnalysis, the AgitationTime selected (`1`) is <2 second or >1 minute, which is outside the suggested range for this experiment: `2`. DFADecaySamplingFrequencyLow In ExperimentDynamicFoamAnalysis, the DecaySamplingFrequency (`1`) selected is low, and would cause data to be sampled only once during the specified DecayTime (`2`) for this experiment: `3`. DFADecayTime In ExperimentDynamicFoamAnalysis, the DecayTime selected (`1`) is <5 second, which is lower than the suggested range for this experiment: `2`. DFANumberOfWashesLow In ExperimentDynamicFoamAnalysis, the NumberOfWashes selected (`1`) is low, and could cause the foam column to be inadequately washed between samples for this experiment: `2`. DigestionAgentsPrecision The DigestionAgents values were provided at higher precision than can be executed in this experiment. The values `1` have been rounded to `2`. DigestionMixRateProfilePrecision The DigestionMixRateProfile values were provided at higher precision than can be executed in this experiment. The values `1` have been rounded to `2`. DigestionTemperatureProfilePrecision The DigestionTemperatureProfile values were provided at higher precision than can be executed in this experiment. The values `1` have been rounded to `2`. DigitalPCRForwardPrimerStockConcentrationAccuracy For sample(s), `1`, the concentration of forward primer identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DigitalPCRMasterMixConcentrationFactorNotInformed For sample(s), `1`, the concentration of MasterMix may be inaccurate as it is calculated using the MasterMixVolume and ReactionVolume. Please inform the concentration factor and leave volume unspecified if possible. DigitalPCRMasterMixQuantityMismatch For sample(s), `1`, the specified MasterMixVolume does not match ReactionVolume/MasterMixConcentrationFactor. Please check that the specified volume is accurate or poor droplet generation and thermocycling will lead to poor quality of results. DigitalPCRMultiplexedTargetQuantity For sample(s), `1`, multiplexing more than 2 targets may result in poor separation of droplet populations and the resulting data may not be useful for target quantification. DigitalPCRProbeStockConcentrationAccuracy For sample(s), `1`, the concentration of probe identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DigitalPCRReferenceForwardPrimerStockConcentrationAccuracy For sample(s), `1`, the concentration of reference forward primer identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DigitalPCRReferenceProbeStockConcentrationAccuracy For sample(s), `1`, the concentration of reference probe identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DigitalPCRReferenceReversePrimerStockConcentrationAccuracy For sample(s), `1`, the concentration of reference reverse primer identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DigitalPCRReversePrimerStockConcentrationAccuracy For sample(s), `1`, the concentration of reverse primer identity oligomer is provided as mass concentration. Inaccuracies in oligomer MolecularWeight will propagate to calculated molar concentration and related quantities. DilutionCurveExcessVolume There will be some leftover dilution volume from samples `3` after making the curve `1` because the instrument only takes `2` volume samples. If you want to keep the leftover dilution set the DilutionStorageCondition option. DiscardedSamples The following object(s) specified are discarded: `1`. DLSDiluentAndBlankBufferDiffer For the following samples, `1`, the Diluent and BlankBuffer options, `2`, are not identical. For B22kD and G22 assays, it is highly recommended that the Diluent and BlankBuffer options are the same, otherwise the resultant B22, kD, and G22 values will not be accurate. Please confirm that the options are specified as desired, or consider letting the BlankBuffer option resolve automatically. DLSDiluentNotSpecified The Diluent option was not specified for the following samples, `1`, and an appropriate Diluent was not able to be determined from the sample's Solvent field. The Diluent for these samples has been set to `2`. For B22kD and G22 assays, it is essential that the Diluent and BlankBuffer options are the same as the input sample's buffer. Please specify the Diluent option for the most accurate experimental results. DLSMoreThanOneCompositionProtein The AssayType is `1` and the following samples, `2`, have Compositions, `3`, with more than one Model[Molecule,Protein]. B22kD and G22 assays should be run on samples which have one Protein in the Composition. For this experiment, the the most concentrated proteins of `2`, `4`, will be used for concentration calculations. Please ensure that the input samples' Compositions are populated correctly. DLSProteinConcentrationHigh The AssayType is B22kD, but the following samples, `1`, have Composition fields with the following Model[Molecule,Protein]s, `2`, with mass concentrations of `3`. The highest recommended concentration of protein for B22kD assays is 25 mg/mL. Please ensure that the Composition fields of the input samples are populated correctly, and that B22kD is the desired AssayType. DNASequencingCartridgeStorageCondition The manufacturer recommends storage of the sequencing cartridge in the refrigerator from 2-8 Celsius, but SequencingCartridgeStorageCondition is not set to Refrigerator or Model[StorageCondition, Refrigerator]. Storing the cartridge outside of these temperatures can degrade the materials in the cartridge and impact performance. It is recommended that the SequencingCartridgeStorageCondition is changed to Refrigerator or Model[StorageCondition, Refrigerator]. DNASequencingInjectionGroupsLengths The specified InjectionGroups have groups of samples that are longer than the NumberOfCapillaries in the SequencingCartridge, which cannot be accommodated by the instrument. The samples have been automatically grouped into InjectionGroups with lengths matching NumberOfCapillaries in the SequencingCartridge. DNASequencingInjectionGroupsOptionsAmbiguous There are replicate samples as input and the samples in are in a different order than the specified InjectionGroups and replicate samples have different options specified, so the matching options to InjectionGroups is ambiguous. All of the sample inputs have been automatically grouped into InjectionGroups with matching options. DNASequencingInjectionGroupsSamplesLengthsMismatch The specified InjectionGroups have more or less samples than are input to the function, or all samples are not specified in a injection group. All of the sample inputs have been automatically grouped into InjectionGroups. DNASequencingInjectionGroupsSettingsMismatch The specified InjectionGroups have values for options DyeSet, ReadLength, PrimeTime, PrimeVoltage, InjectionTime, InjectionVoltage, RampTime, RunVoltage, and/or RunTime that do not match, which cannot be accommodated by the instrument. The samples have been automatically grouped into InjectionGroups with matching options. DNASequencingPCROptionsForPreparedPlate When PreparedPlate->True, PCR is not performed on the samples, so any specified PCR options will be ignored. Please set PreparedPlate to False if PCR is desired. DNASequencingReadLengthNotSpecified The ReadLength has been automatically resolved to 500 base pairs because it could not be automatically determined from the Composition field of the samples `1`. If this value is not correct, please specify a ReadLength value or inform the Composition field in the corresponding sample with a single Model[Molecule,Oligomer] with the PolymerType DNA so ReadLength can be set automatically. DownsamplingRateTooLarge The specified downsampling rate `1` is larger than the half the range of data `2` in dimension `3`. No downsampling will be performed in this dimension. DownsamplingRateTooSmall The specified downsampling rate `1` is smaller than the spacing `2` in dimension `3`. No downsampling will be performed in this dimension. DrainedWell It is expected that the manipulation, `1`, will try and draw more liquid from the location `2` than it will contain at the time (`2` would need an additional `3` to complete the manipulation). DropletAliquotConflict ProbeType is set to Droplet or Surface while Aliquot has been explicitly set to false for samples(s): `1`. Please note that these measurements require removal of a sample for measurement. In the Droplet case you can use RecoupSample->True to collect the sample, but it's not possible to recover the sample used in the surface probe case. DryNoPressure Pressure application was not requested for the following dry `1` sample(s). The lack of pressure may adversely affect measurement, but will continue. DryNoPressureBlanks Pressure application was not requested for the following `1` Blanks. The lack of pressure may adversely affect measurement, but will continue. DuplicateAssignment Multiple assignments in option SplittingAssignments were matched to the same domain(s) `1`. Only the first assignment in this range will be used. DuplicateSamples The samples `1` are duplicated. Therefore the last transport condition will be uploaded and others will be ignored. DuplicateStandardData Multiple sets of standard data were provided for standard analyte(s) `1`. Duplicate standard data will be ignored. DuplicateWavelengths Multiple intensity readings at the requested wavelength were found for `1`. The first reading will be shown. ElisaImproperContainers Polypropylene and Cycloolefine are low protein-binding materials. Choosing these as ELISA plate is likely to affect your results. Consider using a PolyStyrene or other high-protein-binding plate instead. We recommend `2`. ELISANoBlankForExperiment Typically blank is needed for a quantitative ELISA experiment but the current value is Null. If desired, please specify one or more Model[Sample] or Object[Sample] as Blank. ELISANoStandardForExperiment Typically standard is needed for a quantitative ELISA experiment but the current value is Null. If desired, please specify one or more Model[Sample] or Object[Sample] as Standard. EmissionWavelengthOutOfRange The emission wavelength of the following DetectionLabels option at index `1` is outside of the instrument's range: `2`. EmissionWavelength is automatically set to `3` which is the nearest emission wavelength supported by the instrument. Please specify the EmissionWavelength option if a different wavelength is preferred. EmptyCartridgeChannel The selected CartridgeType can test up to `1` analytes but only `2` are specified. To fill up the empty capillary channels, please consider selecting additional supported ExperimentCapillaryELISA pre-loaded cartridge analytes. Please check ExperimentCapillaryELISA help file for the list of current supported analytes. EmptyPostMicellarRegion There are no data points the the provided post-micellar region `1`. In order to fit this region, please change the PostMicellarRange or PostMicellarDomain. To see the surface tensions of the measured samples, you can look in the SurfaceTension field of the data objects. EmptyPreMicellarRegion There are no data points the the provided pre-micellar region `1`. In order to fit this region, please change the PreMicellarRange or PreMicellarDomain. To see the surface tensions of the measured samples, you can look in the SurfaceTension field of the data objects. EmptySplittingDomain The provided domain(s) `1` contain no peaks. Please verify that the PeakGroupDomains option is correct, or adjust peak-picking options to ensure all peaks have been identified. EpitopeBinningAnalysisMissing There is no BindingQuantitation analysis object linked to the input data object(s). If you would like the data to be fit, please run AnalyzeBindingKinetics on the data objects. EpitopeBinningIncompleteDataSet The data objects do not test every possible pairing. The binning analysis cannot be conducted, only the raw response value for each pairing will be given in the output. EpitopeBinningLargeFitDomain The FitDomain value `1` exceeds 20% of the length of the average competition data set. This may impact the validity of the results. EpitopeBinningSpeciesNotFound The following objects used as slow binding species were not found in the inputs or the composition or models of the inputs: `1`. ESITripleQuadTooManyCalibrants For this protocol running in ESI-QQQ, there are `1` calibrants are not the default calibrants of this instrument: `2`. For the precision of the the instrument, only the first one,`3`, will be used for the calibration: EvaporateTemperatureGreaterThanSampleBoilingPoint In ExperimentEvaporate, the specified EvaporationTemperature for `1` is greater than the sample's BoilingPoint, `2`, and may result in sample degradation. An EvaporationTemperature below the SolventBoilngPoint is recommended. ExcessDestinationVolume The primitive at index `1` has Destination volume of `2` that exceeds the specified FinalVolume. ExcessiveCheese While nutritional science often suffers from a lack of large long-term double-blind randomized controlled trials, the data may suggest that choosing to consume such a large amount of cheese could be hazardous to your health. You may wish to consider specifying a lower CheeseWeight. ExcessiveSolventVolume The SolventVolume `1` is greater than the LoadingSampleVolume `2` for the following input sample `3`. Extra LoadingSample solution will be stored under the storage condition of the Solvent option. To avoid extra solution storage, consider setting SolventVolume to Automatic or to a volume equals to the LoadingSampleVolume. ExcitationWavelengthOutOfRange The excitation wavelength of the following DetectionLabels option at index `1` is outside of the instrument's range: `2`. ExcitationWavelength is automatically set to `3` which is the nearest excitation wavelength supported by the instrument. Please specify the ExcitationWavelength option if a different wavelength is preferred. ExcludeOverwriteInclude Exclude contains point also in Include. Any peak whose range contains this point will be excluded from the returned peaks. ExistingModelReplacesInput An existing model (`1`) fulfills the input template model (`2`) with specified options. The existing model (`1`) will be used as the alternative preparation template for this stock solution. ExistingPipettingMethod The model(s) `1` already have the default pipetting method(s) `2`. This existing method will be overwritten. ExpiredSamples The following sample objects are marked expired: `1`. ExplicitNullOptions Options `1` cannot be explicitly specified as Null. Null values will be ignored, and these options have been set to their default values. ExtCoeffNotFound The ExtinctionCoefficients field is not populated for the model(s) of the following input sample(s): `1`. Please upload ExtinctionCoefficients to the corresponding Model in the form of {<Wavelength->_, ExtinctionCoefficient->_>}. Defaulting wavelength to 260 nm (if not a protein or peptide) or 280 nm (if a protein or peptide). Extrapolation Input datasets at indices `1` contain values outside of the range of provided standard data. Extrapolation will occur when applying standard curves. FailedToComplete Simulation failed to run to specified "simulationDuration" completion time. Try increasing AccuracyGoal and PrecisionGoal, and decreasing MaxStepFraction. FewerLaddersThanUniqueOptionSets Fewer ladders than the number of unique sample preparation and separation conditions applied to Samples (`1`) were specified. Only the most common set of conditions will be applied to the specified ladders. Please consider specifying exactly `1` instance(s) of each unique ladders in order to apply all unique sets of options on these ladders. FillToVolumePrimitiveIncluded Usage volume calculation for FillToVolume primitives may not be accurate due to possible density changes during manipulation. FillToVolumeSolventDefaulted The Solvent option could not be automatically set for the following input sample(s) or container(s) because the option was not specified and the Solvent field is not sufficiently populated: `1`. This option has been set to Model[Sample, "Milli-Q water"], but if this is not satisfactory please specify the Solvent option directly. FilteredAnalytes The given analytes, `2`, will be filtered to only include, `3`, since the given analytes occupy a large mass range that will lead to a poor resolution for the given instrument. Please only include analytes that occupy a similar mass range or enqueue multiple runs for the sample(s), `1`, with different mass ranges. FirstWavelengthNotUsedForAdaptive The following AdaptiveExcitationWaveLength option for pool number `1` does not match the first ExcitationWavelength which will be used to acquire images in each sample pool: `2`. For pool number `1`, choosing the following excitation wavelengths will provide the best results for adaptive sampling: `3`. FitIssue The NonlinearModelFit produced errors in fitting a logistic function to the curve `1`. "Gradient" method will be passed to NonLinearModelFit to improve the fit results. Please consider increasing the number of data points in the experiment by changing the temperature resolution. FlashFreezeAliquotSampleStorageWarnings In ExperimentFlashFreeze, the AliquotSampleStorageCondition (`1`) is not CryogenicStorage for the following samples, which may result in storage at insufficiently cold temperatures: `2`. FlashFreezeNoVolume In ExperimentFlashFreeze, the following samples `1` do not have their Volume field populated. The Volume of the sample must be known in order to determine if the sample can be safely flash frozen. FlashFreezeSampleInStorageWarnings In ExperimentFlashFreeze, the SamplesInStorageCondition (`1`) is not CryogenicStorage for the following samples, which may result in storage at insufficiently cold temperatures: `2`. FlowCellChangedToNearest The specified flow cell pathlength `1` is not available for the instrument (the following are `2`). The flow cell pathlength was set to the nearest available one at `3`. FluidAnalysisMeasurementMismatch The option FluidAnalysisMeasurement for the primitives at index `1` is set to a value not intended for the composition of samples `2`. The option can be left as is or if desired, can be left to be set automatically according to the sample's composition. FluidTypeCalibrationMismatch The option FluidTypeCalibration for the primitives at index `1` is set to a value not intended for the composition of samples `2`. The option can be left as is or if desired, can be left to be set automatically according to the sample's composition. FormulaOptionsUnused The options `1` have been specifically set; however, these options only apply when a new stock solution formula is provided as input in order to upload and prepare a new stock solution model. Since existing stock solution models were provided as input, these options will be unused. FreezeCellsIgnoredFreezerObject Currently, ExperimentFreezeCells does not support specification of freezer locations. Specified freezer instruments `1` will be ignored, and the location selection will be handled by the storage system based on the specified instruments. FreezeCellsIgnoredOptions The specified options `1` are not consistent with the specified method for batches at indices `2`. These options will be ignored. Please specify a different method if you wish to utilize these options. FreezeCellsInstrumentalPrecision The specified precision for option(s) `1` is beyond the capabilities of the instrument. The values of these options will be rounded from `2` to `3`. FreezeCellsInvalidTransportConditions The specified TransportConditions are not set to Minus40 or Minus 80 at indices `1`. Please ensure that these conditions are acceptable for the transport of the frozen cells. FreezeCellsLowMeltingCoolant Coolants `1` have melting points that are below the specified or default FreezingCondition. Upon freezing, the coolants may lose efficiency in heat transfer, and result in unpredictable cooling rates. Please ensure that the specified coolants are acceptable for this experiment. FreezeCellsMixedMethodTypes FreezingMethods contain more than one type of method without specification of Batches. Samples will be distributed between InsulatedCooler and ControlledRateFreezer batches to minimize the number of batches, and spread samples as evenly as possible between batches. Please specify Batches if you would like to control how samples are processed. FreezeCellsProfileContainsWarmingStep Specified FreezingProfiles contains a warming step at indices `1`. Please ensure that this step will not cause any issues for the freezing process. FreezeCellsWarmingDuringTransport The specified TransportConditions is above the final freezing temperature of the cells at indices `1` and will cause warming of the cells during transport. Please ensure that the warming during transport will not cause problems with the cells. FreezeCellsWarmStorageCondition Specified StorageConditions for samples `1` are warmer than the final freezing temperature of the samples. Please ensure that the increase in temperature during final storage is acceptable for this experiment. GCColumnMinTemperatureExceeded Temperature setpoint(s) `1` in option `2` are below the minimum column temperature of `3`! Your column may not be able to perform separations at this temperature. GradientAmbiguity Full gradients were defined for the options: `1` which conflict with the specified, adjacent GradientA/B/C/D options. Consider only defining `1` and not the corresponding GradientA/B/C/D options. GradientNotReequilibrated The gradients occurring before the sample injections, do not reequilibrate the gradient. This may lead to the analyte prematurely eluting. Consider setting ColumnRefreshFrequency -> 1. GradientShortcutAmbiguity Gradient shortcut options are specified in both `1` and `2` A/B/C/D options but they conflict with each other. The experiment will commence as directed with the latter value(s). Please verify the resolved gradients in the submitted protocol are desired. HighFlowRate In ExperimentEvaporate, the FlowRate is high and may result in sample splashing and cross-contamination for the following samples: `1`. A lower flow rate is recommended. HighRamanSpectraDensity The number of spectra that are plotted may make the output difficult to interpret and increase the amount of time it takes for plots to generate, set ReduceData ->True or Automatic to reduce the data density to 10%. HighWellVolume After injections have completed the following samples will have volumes over 250 microliter: `1`. As the well volume approaches the maximum, sample can splash onto the optics. Consider using the lowest injection flow rate and/or lowering the involved volumes. HomogenizationAmplitude The homogenization amplitude for the sample(s), `1`, are set to over 50%. Depending on sample viscosity, this can cause the sample to be forcibly displaced outside of its container. This can be safeguarded against by making sure that the sample's volume is not more than 50% of the container's MaxVolume. HPICGradientShortcutAmbiguity Gradient shortcut options are specified in both `1` and `2` options but they conflict with each other. The experiment will commence as directed with the latter value(s). Please verify the resolved gradients in the submitted protocol are desired. HPLCBufferConflict The resolved values of `1` differ from provided gradient method(s). The experiment will commence as directed with the former value(s). HPLCFluorescenceSamplingRateAdjusted The following fluorescence sampling rate options, `1`, can only be set values (`2`) and were set to the closest one. HPLCGradientNotReequilibrated The gradients occurring before the sample injections of `1` at index `2` of the input samples, do not reequilibrate the gradient. This may lead to the analyte prematurely eluting. Consider setting ColumnRefreshFrequency -> 1. HPLCpHCalibrationBufferSwapped The LowpHCalibrationTarget is larger than HighpHCalibrationTarget. The two calibration buffers will be swapped during calibration. Please choose different calibration buffers if this is not desired. ImagingIncompatibleContainer The sample(s) `1` are in containers that are not compatible with any available imaging apparatus. Samples will be transferred to new containers to allow imaging to occur. IncompatibleAnalytes `1` type analytes are not recommended for this assay. Automatic options will default to oligomer settings if no other compatible analytes are present. IncompatibleCalibrant The molecular weights of the analytes (either provided by the user with the option Analytes or deduced from the sample's composition) in the sample(s), `1`, fall outside the range formed by the reference peaks in the calibrants. Consider choosing a calibrant with reference peaks which would flank the expected sample peak(s). IncompatibleColumnAnalysisChannel The specified `1` `2` has a preferred analysis channel of `3`, which is different from the specified `4` AnalysisChannel. Please consider specifying a different `1`. IncompatibleColumnTechnique The specified `1` `2` has a ChromatographyType of `3` and may not perform as expected using the `4` ChromatographyType. IncompatibleColumnTemperature The specified `1`s (`2`) are outside the specified `3` (`4`) temperature range of `5` to `6`. To prevent column damage, use column temperatures within this compatible range or specify a different column. IncompatibleColumnType The specified `1` `2` has a SeparationMode of `3` which is not the same as the specified SeparationMode (`4`). IncompatibleICDetector The specified detectors `1` are not a subset of compatible detectors `2` for the specified analysis channel `3`. Data will only be collected on the compatible detectors. IncompatibleMaterials The provided samples, `1`, are chemically incompatible with the wetted materials in `2`. Please check the IncompatibleMaterials listing in these chemicals, and the WettedMaterials of the instrument, and substitute these samples for chemically compatible ones if possible. IncompatibleSliceUnits Incompatible units `1` provided in DataSlice option; expected units compatible with `2` for dimension `3`. Units will be ignored. IncompatibleTransformation A 3D transformation `1` is selected for a 2D DataSet `2`. The transformation is switched to None. IncompatibleTrimmingOptions The trimming options `1` are incompatible with the TrimmingMethod `2` and will be ignored. Please remove these options, or change the TrimmingMethod. IncompatibleUnits The option value `1` for `2` is incompatible with resolved input unit `3` in `4`. Units will be ignored for this dimension. InconsistentOptionName The input option name `1` and option OptionName `2` are not consistent. The input option name will be used. IndexMatchingConflicts Options index matched to `1` must all be the same length. Please check your provided option values for: `2`. Their current lengths are: `3` IndexMatchingOptionMissing Options index-matched to `1` cannot have their lengths fully validated because this option was not provided. The following options will be assumed to have valid lengths: `2` InitialConditionPadding Padding non-species-specific initial conditions `1` with zeros to `2` so it matches the number of model species `3`. InputContainsTemporalLinks The following input(s), `1`, were detected to be links. The most recent information about these inputs will be downloaded in order to generate an protocol and the temporal date provided will be ignored. InputFieldOverride The input field(s) for input(s) `1` (`2`) do not match the option(s) InputField, and have been defaulted to the option value(s) `3`. InputLengthMismatch The number of concentrations (`1`) does not match the number of species (`2`). Please ensure that both inputs are of the same length. Padding both inputs to match the greater of the two lengths. InputTooLong Input can be a maximum of two lists of oligomers. Truncating input to first two lists. InstrumentPrecision The machine precision of `1` has a resolution of `2` and therefore `3` will have to be rounded from `4` to `5` to proceed. If you don't wish rounding to occur automatically, please supply a value within the given resolution. InstrumentUndergoingMaintenance The following specified instruments are currently undergoing maintenance: `1`. This protocol may still be confirmed, but will not be run until these instruments are no longer undergoing maintenance. InsufficientAmount The amount of `1` is not sufficient to perform the manipulation at index `2` and all subsequent manipulations. Please check the sample in question, or provide a new sample with sufficient amount. InsufficientBlankSpace There are not enough empty wells in the assay container to transfer in the blank samples so everything must be aliquoted into a new container. InsufficientBLIStartDelay The current StartDelay `1` may be insufficient for the assay plate to reach the desired temperature `2`. For a non-ambient temperature a 10 minute StartDelay is recommended. InsufficientPreparedMicrowaveDigestionSample Total of SampleAmount and DigestionAgents is not above the minimum of `1` for all samples. The microwave reactor will attempt to achieve the desired temperature program, but may not heat and/or stir uniformly, leading to hot or cold spots within the sample mixture. The temperature data produced may no longer be representative of the reaction conditions. InsufficientSampleVolume The samples `1` do not have sufficient volume for the injection volume (default to 10 uL for Analytical measurement or 500 uL for Preparative measurement) plus the autosampler's dead volume of `2`. The experiment will still attempt to make injections with what is currently available; please change the InjectionVolume options if this is not desired. InsufficientSDSinSample The final concentration of SDS in the the samples `1` is lower than the recommended 0.5% SDS minimum. Please consider adding more SDSBuffer to your sample or use a more concentrated SDSbuffer. InsufficientStaticDialysateVolume In order for the stir bar to move unimpeded, we recomemnd a dialysate volume of at least 1 Liter. InsufficientVolume The experiment requests more volume/mass/count of `1` (`2`) than is currently available (`3`). Please check the available volume of these sample(s), or provide an alternative sample or model. InvalidAmountLength The amount(s) specified in the primitive at index `1` have invalid lengths that do not match with Sources or Destinations. Please check the definition of the primitive in question and specify amounts with correct lengths InvalidCompositionUpdate The Composition fields of SamplesIn could not be updated because computed concentrations have units of `1`, which do not match CompositionP. UpdateSampleComposition does not support non-standard concentration units; if updating is desired, please ensure that each entry in StandardConcentrations matches one of the supported units in CompositionP. InvalidDuration The data packet (`1`) was collected over `3`, while `2` was specified as Duration. InvalidFluorescneceExposureTime The data packet (`1`) does not contain data collected at the specified FluorescenceExposureTime. Data is present for the following exposures `2`. Defaulting to plot data at the longest exposure present (`3`). InvalidFramesToPlot The data packet in index (`1`) does not contain data collected for frame `2` (data is present for `3` frames). Please make sure to specify a valid value. Defaulting FramesToPlot option to All. InvalidLaserOptimization `1` option(s) will be ignored. Laser power cannot be adjusted on `2` instrument. InvalidNameReference The names `1` used in primitives at index `2` do not have an associated Define primitive that establishes the name. Please define any used names using a Define primitive. InvalidNumberOfSpecies `1` species were requested but only `2` are present in the input data. Please check that the requested number is correct. Defaulting to n=`2`. InvalidOverlayChannels At least one specified channel for the ChannelsOverlaid option of object(s) `1` is invalid. Please only include channels that are present in the given data object(s). Attempting to use any available specified channels. InvalidPolymerType Polymer->Null is not a valid option value for vague input in function `1`. Polymer will be defaulted to Automatic. InvalidPosition There is no fraction at position `1`. Only `2` fractions were collected. InvalidPrimitiveKeyValue The value of the following key(s) defined in the primitive at index `1` have invalid pattern that may affect the usage amount calculation: `2`. Please check the definition of the primitive in question. InvalidPrimitiveType The primitive(s) at index `1` does not have the type Transfer, Aliquot, Consolidation, Resuspend, FillToVolume, or Define. The usage amount will only be calculated from primitives with valid manipulation type. InvalidRangeSpecification The provided plot range `1` is invalid and has been defaulted to Automatic. A valid plot range has the form {{xmin,xmax},{ymin,ymax}}. Each {min,max} may also be specified as All, Full, or Automatic. InvalidSampleAnalyteConcentrations The analyte concentration could not be determined for the following samples: `1`. Without the Analyte concentration, the resulting sample composition cannot be accurately determined. Please update the sample object Analyte field and ensure that all MolecularWeights are informed. InvalidStandardConcentrationLength The length of the StandardConcentrations option (`1`) does not match the length of the Standards field of the input protocol (`2`). The StandardConcentration option will be ignored, and standard concentrations will be computed from the Composition fields of Standards in the input protocol. InvalidStandardDataLength The length of the StandardData option (`1`) does not match the length of the Standards field of the input protocol (`2`). StandardData will be defaulted to the StandardData field from the input protocol, and the StandardData option will be ignored. InvalidStandardDataReferences One or more of the SamplesIn fields of objects in the StandardData option do not match the Standards field of the input protocol. StandardData will be defaulted to information from the input protocol, and the StandardData option will be ignored. InvalidStandardsNoPeaks The standard(s) at indices, `1`, do not have molecules assigned to their peaks in their latest peaks analysis object, and will be ignored in composition analysis. Please add these indices to the ExcludeStandards option, or let the ExcludeStandards option resolve automatically. InvalidTargetUnits The target units specified `1` do not match the option TransformX. Plot will be generated without the transform. InvalidTrimRange The provided TrimAfterIndex (`2`) must be greater than or equal to the provided TrimBeforeIndex (`1`). Please specify a valid trimming range. InvalidWindowSize The provided WindowSize (`1`) must be 2 or greater and less than the length of the sequence. Please specify a WindowSize in the range {2,`2`}. ItemsContainerless The DiscardContainer option will be ignored for the items `1`, which do not have containers. DiscardContainer will only have effect for samples or containers. JobNotFound The job(s) `1` could not be found in Constellation. Please ensure these objects exist, and that you have permissions to access them. LabelFieldInvalid The specified LabelField (`1`) does not contain a list of labels. Please check that the field name is correct, or please consider using the ContourLabels option to set the contour labels directly. Defaulting LabelField to None. LabelFieldLengthMismatch The specified LabelField (`1`) contains a list of labels whose length does not match the 3D input data. Please check that the field name is correct, or please consider using the ContourLabels option to set the contour labels directly. Defaulting LabelField to None. LabelFieldNotFound The specified LabelField (`1`) was not found within the input data type (`2`). Please check that the field name is correct, and please see the Labels option for specifying manual contour labels. Defaulting to using the contour indices as labels. LabelsLengthMismatch Length of CountourLabels (`1`) does not match the number of contours (`2`). The contour z-coordinate values will be used as labels instead. If custom labels are desired, please try adding or removing some labels until the lengths match. LargeCellSampleBatch The batches, `1`, contain more than 2 cell samples. Large batches can result in cells being exposed outside of the incubator for longer than ideal times. It is recommended to process no more than 2 cell samples in a batch (cells will be placed in the incubator before and after batches are processed). LCMSAutoResolvedNeutralLoss For using TripleQuadrupole and NeutralIonLoss as the acquisition mode, a valid mass value in NeutralLoss option needs to be specified in order to generate useful and reliable data. The experiment can further procees with an automatically assigned value `2` for the following options `1`, but this experiment may not generate reliable data. LimitedReferencePeaks In order to calibrate the instrument it is recommended that there be at least 3 calibrant peaks in the specified mass ranges. Please consider adjusting the following mass ranges: `1`. LocalAsynchronousNotSupported Asynchronous computing is not supported for local computations. Please set Computation->Cloud for asynchronous computations. Compute will proceed with WaitForComputation->True. LongComputation This computation is estimated to take `1` to complete, with an estimated completion time of `2`. LongLeadTimeCartridge The cartridge is a pre-loaded cartridge that is not in stock at ECL and may take 14 days to arrive. To start the experiment sooner, please consider testing a total of fewer than 48 samples and selecting Model[Container,Plate,Irregular,"Human 48-Digoxigenin Cartridge"] as the Cartridge. LongRamanSamplingTime The measurement time for samples `1` exceed 20 Minutes. Consider adjusting the parameters `2` to reduce the resolution, duration, overlap, or size of the sampling pattern. LowDLSReplicateConcentrations The NumberOfReplicateConcentrations is set to 2. It is recommended to have 3 or more replicates per dilution curve concentration for B22/kD and G22 assays. LowMinMass For input samples `1`, the instrument `2` is likely to produce noisy data in the lower mass range requested because a high abundance of background peaks in this mass range is common. Consider adjusting the lower limit of the MassDetection to above `3`. LowSurfaceTensions The samples, `1`, have low surface tensions `2`. The should ManyTransfersRequired Some of the specified manipulations contain volumes >10mL which will require many discrete pipette transfers to complete on the liquid handler. You may wish to consider performing some or all of your manipulations in a separate MacroLiquidHandling protocol. MappedNullPrimaryData No primary data was found in field(s) `1` of `2`. Please check your inputs. MassKnown For samples `1`, the MeasureTotalWeight is set to True although the Mass of the sample(s) is known. As a result the total weight of the sample(s) will be re-measured and the Mass in Object[Sample] overwritten with the newly recorded value. Consider leaving MeasureTotalWeight blank or set it to False, if you would like to calculate the count using the currently recorded Mass. Measure620WithCorrection 620 Nanometer is specified in AbsorbanceWavelength or PrereadAbsorbanceWavelength while SignalCorrection is True. Note that, if continue, the correction wavelength (which is is always 620 Nanometers) will be turned off when reading an AbsorbanceWavelength or PrereadAbsorbanceWavelength at 620 Nanometers. Other wavelengths, if specified, will not be affected. MeasurementMayRequireTransfer Please be aware that these models, `1`, may be transferred to another container in order to experimentally determine the amount. If this is not desired, please specify an amount or specify ProductDocumentation to indicate that the amount should be found on the documents that ship with the sample. MeltingPointBadData The `1` `2` does not have the expected shape. The resulting melting point may be inaccurate. MeltingPointOutOfDomain Warning: MeltingPoint is out of input Domain. MicroscopeImageSelectNoData There is no data provided in the following protocol(s): `1`. MicrowaveDigestionMissingRecommendedVenting When the SampleType is Inorganic or Tablet, PressureVenting is recommended for `1`. No reaction venting can lead to pressure buildup, reaction vessel failure, and premature reaction shutdown. MicrowaveDigestionPossiblePreparedSample The samples `1` contain standard digestion agents in the Composition field. To dismiss this warning, set PreparedSample to True or False. MicrowaveDigestionSampleAmountPrecision The SamplesAmount values were provided at higher precision than can be executed in this experiment. The values `1` have been rounded to `2`. MicrowaveDigestionUncrushedTablet It is recommended that CrushTablets be set to True for all tablet inputs. The sample(s) `1` will not be crushed and may not digest completely. MismatchedColumnAndScale The amount of active sites `1` available on `2` disagrees with the specified scale `3`. Be advised that options that automatically resolve based on scale may not be optimal for the amount of resin used. The experiment will proceed with the specified options. MismatchedCoverslipThickness The following specified CoverslipThickness options for samples in pool number `1` do not match the value in their container models: `2`. No action is needed if this is intentional. MismatchedEmissionWavelength The specified EmissionWavelength option `1` at index `2` does not match the emission wavelength of the following molecules in DetectionLabels option: `3`. No action is needed if this is intentional. MismatchedExcitationWavelength The specified ExcitationWavelength option `1` at index `2` does not match the excitation wavelength of the following molecules in DetectionLabels option: `3`. No action is needed if this is intentional. MismatchedPeakCounts Multiplicities `1` at indices `2` in option PeakGroupMultiplicity are inconsistent with the number of peaks in the corresponding splitting groups `3`. J-coupling constants will not be calculated for these groups. MismatchedPlateBottomThickness The following specified PlateBottomThickness options for samples in pool number `1` do not match the value calculated from Dimensions, WellDepth, and DepthMargin of the container model: `2`. No action is needed if this is intentional. MismatchedTemplateOptions The template protocol contains options which not match the length of your provided options/inputs. These options cannot be used and will be set to their default values. If you don't wish to use the defaults, please directly specify new values for these options. Check: `1` MismatchingOptionsWarning `1` for the following input samples `3`. Please `2`. MissingBLIProbeEquilibrationOptions When ProbeRackEquilibration -> True, the following options must be set to Automatic or a non Null value: `1`. MissingCleaningMethod The CleaningMethod for the probes `1` is Null, while SingleSamplePerProbe is `2`. This may result in contamination of the samples as the probes will not be cleaned between measurements. MissingDilutionCurve A Diluent `1` is specified, but no DilutionCurve is given `2` for the samples `3`. MissingDilutionMixRate A DilutionCurve is specified `1`, but no DilutionMixRate is given `2` for the samples `3`. The dilution curve will be prepared without mixing. MissingDilutionMixVolume A DilutionCurve is specified `1`, but no DilutionMixVolume is given `2` for the samples `3`. The dilution curve will be prepared without mixing. MissingDilutionNumberOfMixes A DilutionCurve is specified `1`, but no DilutionNumberOfMixes is given `2` for the samples `3`. The dilution curve will be prepared without mixing. MissingObjects Unable to find object(s) `1` in the database. Please check for errors in the object IDs or names. MissingSampleComposition Sample composition is missing for `1`. As a result, sample Volume can't be accurately calculated to reach 1 mg/mL protein and defaulted to 25% of the TotalVolume. Please specify the volume if desired. MissingVolumeInformation The sample(s) `1` are missing volume information. The MaxVolume of the source containers will be used instead when resolving Automatic options. MixedBasesUnsupported The requested Method `1` does not support assignment of MixedBases. Please set MixedBases to False, or choose a Method that supports mixed base assignments (`2`). MixedDataTypes You are trying to plot data with types `1`. Only one type of data can be plotted at a time. MixerChangedToNearest The specified mixer volume `1` is not available for the instrument (the following are `2`). The flow cell pathlength was set to the nearest available one at `3`. MixVolumeGreaterThanAvailable The sample(s), `1`, do not have enough volume to be mixed by pipette with the following MixVolume(s) specified, `2`. When being mixed, these samples will be overaspirated. If you do not want this to happen, please change the value of the MixVolume options. MoatAliquotsRequired Note that in order to create a moat samples will be aliquoted into a new plate. ModelDensityNotUpdated The Models `1` already have Density information and will not have their Density updated during the course of this protocol. ModelDensityOverwritten The Models `1` already have Density information, which will be overwritten as the result of this protocol. ModelNotLiquid Specified model(s) `1` do not have a Liquid State. Pipetting parameters specified may not perform as specified when manipulating non-liquid models. ModelVolumeOverload As neither the BedVolume nor the ResinLoadingAmount has been specified, the BedVolume has been set to the BedVolume provided in the packed Column Model. The ResinLoadingAmount will be backcalculated from the BedVolume and the value of the SpecificVolume for the set resin and storage buffers, `1`: MolecularAssignment Unable to automatically assign peaks to any of the nuclei in KnownSpecies `1`. Please ensure peaks have been correctly labeled, or manually provide assignments using option SplittingAssignments. MolecularWeightNotFound Could not download field MolecularWeight from `1`. Please use Inspect to verify that MolecularWeight is defined for these inputs. MoleculeInfoNotFound Could not download a valid Molecule or MolecularFormula from `1`. Please use Inspect[] to verify that at least one of these fields are present in object(s). MoreThanOneTypeOfAnalyte The input samples contain `1` analytes. More than one type of analyte is not recommended for this assay. Automatic options will default to oligomer settings. MultipleAnalytes For the following samples `1`, there are multiple Analytes. The first member of Analytes will be used. MultipleAssayPlates Samples `1` with no dilution `2`, `3` are specified with a True SingleSamplePerProbe `4`. Since there are more than 8 samples and the samples are not sharing probes, this will take multiple assay plates. If you would like the samples to share probes and all be on the same plate, set SingleSamplePerProbe to False. MultipleElectrochemicalDetectionModes The specified waveforms in either `1` option or the injection table have different `2`. Please consider specifying waveforms with the same detection mode. MultipleEmissionWavelengths The following molecules specified as DetectionLabels option at index `1` do not share a single emission wavelength: `2`. EmissionWavelength is automatically set to `3` based on the average of all molecules' emission wavelengths. Please specify the EmissionWavelength option if a different wavelength is preferred. MultipleExcitationWavelengths The following molecules specified as DetectionLabels option at index `1` do not share a single excitation wavelength: `2`. ExcitationWavelength is automatically set to `3` based on the average of all molecules' excitation wavelengths. Please specify the ExcitationWavelength option if a different wavelength is preferred. MultipleIncubationTimes The sample(s), `1`, have multiple Time option(s) specified. For these sample(s), only the maximum of the Time option(s) specified (`2`) will be used for all sample(s). If this is not desired, please split the sample(s) with different desired incubation time(s) into separate protocol(s). MultipleSampleOligomersSpecified The composition of the sample(s) `1` contains multiple oligomers of different lengths; consequently, the ReadLength option is automatically resolved to 500 base pairs. Please specify a ReadLength or use a sample containing a single oligomer if this value is not correct. MultipleTargets For the following analytes `1`, there are multiple Targets. The first member of Targets will be used. NeedConcentrationForMolarEllipticity For samples `1`, if calculating the molar ellipticity at every detection wavelenght is desired, valid AnalyteConcenctions needs to be specified for the sample, the concentration `2` cannot be used to calculate molar ellipticity. NegativeConcentration Simulation produced negative concentrations which indicates an inaccurate result. Try increasing AccuracyGoal and PrecisionGoal, and decreasing MaxStepFraction. NegativeDiluentVolume A diluent volume is calculated to be negative in your sample or standard dilution. This is typically cause by the total volumes of concentrated spike, primary antibody, or capture antibody exceeding the diluent volume specified in your dilution curves. Note that the volumes of these additives to your samples count towards the volume of diluent. If you think these volumes are too small to make a difference in the total volume, you can ignore this message and the diluent volume will be rounded up to 0 Microliters and your specified volumes of spike, primary antibodies, and capture antibodies to be mixed with samples will not be changed. NephelometryIncomputableConcentration For sample(s) `1`, the input analyte concentration cannot be calculated because the analyte(s) `2` do not have `3` fields populated. NestedIndexLengthMismatch Nested index-matched option/s `1` are not matching in length for samples `2`. Defaulting to expand with Automatics or trim trailing values based on the `3`. Please make sure input lengths match in order to avoid unintended modifications to your method. NewModelCreation A new model (`3`) will be created when using `2` as input template model, because the options `1` are different from the ones in the input template model `2`. NewServiceProviderForModel The specified service providers, `2`, are not known for these models, `1`. The model will be updated with this service provider. No action is needed on your part. NoAlkylatingAgentIdentified A alkylating agent was not identified in the composition of the alkylating agent object specified for `1`. Please make sure that the object has valid composition or specify the alkylating agent's volume. NoAnalyteInAnalysisChannel There is no analyte currently in the `1` while Standard or Blank samples are specified. NoAnalytesFoundInStandards The analyte `1` could not be matched to a nonzero concentration in the Composition of any of the Standards from the input protocol. Please include a standard which contains a non-zero amount of this analyte. NoBindingKineticsAssociationBaseline No baseline could be identified in `1`. If a baseline is desired, specify one using the AssociationBaseline option. NoBindingKineticsDissociationBaseline No baseline could be identified in `1`. If a baseline is desired, specify one using the DissociationBaseline option. NoBLIProbeEquilibration The probes are not equilibrated prior to use, which will likely adversely impact their performance. Set ProbeRackEquilibration to True, to equilibrate probes in the probe rack. The lack of a probe rack buffer can lead to mis-calibration of the bio-probes which may result in loss of data or inaccurate results. NoBubblesFound No bubbles could be identified in the video data associated with input `1`. Please ensure the video is in the correct format, and that the MaximumBubbleRadius and MinimumBubbleRadius options are set to reasonable values. NoData There is no data provided in the given protocol(s) `1`. NoDataInChannel No data was found in channels `2` of input `1`. These channels will be ignored in the ClusteredDimensions option. NoDataInDomainRange No data points remain after applying Domain and Range filtering. NoELISAAssayTypeForAntibodySamples ELISA is not a member of AssayTypes for the antibody samples `1`. The sample may have not been evaluated/certified for ELISA application by the manufacturer and the experiment may not give optimal results. Please check to make sure desired antibody samples are selected. NoMixingDespiteAliquotting For the following sample(s) or sample pool(s), `1`, PooledMix was set to False although pooling or aliquotting into a cuvette is taking place. Mixing samples after aliquotting is recommended to ensure a homogeneous solution for the absorbance measurement. If you would like to mix these samples after aliquotting, turn PooledMix to True or leave it Automatic. NonAnhydrousSample The following liquid samples, `1`, at manipulation indices, `2`, are not marked as Anhydrous->True. Please verify that these samples will not invalidate the atmosphere of the glove box. Any damage that is done to the glove box will be billed to the user and could result in the loss of ECL privileges. NonAscendingOrderedDataSet {x,y} coordinates of the dataset `1` is not in monotonically increasing order. This dataset will be sorted for smoothing purposes. If you intend to not to use sorting, select AscendingOrder-> False. NonDefaultDLSSamplePreparationPlate The specified SamplePreparationPlate, `1`, is of `2`. The default SamplePreparationPlate Model is Model[Container, Plate, "96-well PCR Plate"]. Using a non-default Model, particularly one with a larger MinVolume, may result in mis-loading of the AssayContainers, as the small 9uL transfers have been optimized only for the default model. Please consider using the default SamplePreparationPlate Model. NonDefaultPolishingSolutions The following solution(s) `1` provided in the PolishingSolutions does/do not match with the default polishing solution model in the corresponding polishing pads, for the input sample `2`. Please consider to set PolishingSolutions for this sample to Automatic or provide a list of default polishing solutions models in the polishing pads. NonDefaultTotalProteinQuantificationReaction The QuantificationReactionTime, `1`, and QuantificationReactionTemperature, `2`, have been specified, but the AssayType, `3`, is not BCA. Bradford and FluorescenceQuantification assays should have the absorbance or fluorescence values read shortly after mixing the samples and the QuantificationReagent. Please double-check that these options have been specified correctly. NonEmptyContainerNotAutoclaveSafe The following containers, `1`, are not autoclave-safe and have non-Solid Contents. We will attempt to move the contents into the following autoclave-safe containers, `2`. If this is not desired, please choose a different input container or transfer the contents to a desired container. NonEqualSpacedDataSet {x,y} coordinates of the dataset `1` is not equally (evenly) spaced. An equally spaced dataset will be generated for smoothing purposes. If you intend to not use equal spacing, select EqualSpacing-> False. NonIdealWesternBlockingBuffer The following BlockingBuffers, `1`, are not the ideal BlockingBuffers for the Organism of the PrimaryAntibody of the following pairs of input samples and antibodies, `2`. Please consider letting the BlockingBuffer option automatically resolve. NonIdealWesternPrimaryAntibodyDiluent The following user-supplied PrimaryAntibodyDiluents, `1`, are not the ideal PrimaryAntibodyDiluents for the Organism of the PrimaryAntibody of the following pairs of input samples and antibodies, `2`. Please consider letting the PrimaryAntibodyDiluent option automatically resolve. NonIdealWesternSecondaryAntibody The following SecondaryAntibodies, `1`, are not the ideal SecondaryAntibodies for the Organism of the PrimaryAntibody of the following pairs of input samples and antibodies, `2`. Please consider letting the SecondaryAntibody option automatically resolve. NonIdealWesternStandardPrimaryAntibody The following StandardPrimaryAntibodies, `1`, are not the ideal StandardPrimaryAntibodies for the Organism of the PrimaryAntibody of the following pairs of input samples and antibodies, `2`. Please consider letting the StandardPrimaryAntibody option automatically resolve. NonIdealWesternStandardSecondaryAntibody The following StandardSecondaryAntibodies, `1`, are not the ideal StandardSecondaryAntibodies for the StandardPrimaryAntibody and the Organism of the PrimaryAntibody of the following pairs of input samples and antibodies, `2`. Please consider letting the StandardSecondaryAntibody option automatically resolve. NonMappableOption Please check option `1`. You can specify a single value which will be applied to all plots. Alternatively you can specify a list with one value per plot. The default value will be used for this option. NonNumericLabelField The specified LabelField (`1`) does not contain a number or quantity for one or more of the input data objects. If ContourSpacing is set to Value, the values obtained from LabelField will not be used to set each contour's Z-coordinate. They will still be used to set the value of ContourLabels if ContourLabels is set to Automatic. NonOptimalCapillaryELISASampleDilution It is recommended to dilute the samples `1` to at least the minimum dilution factor `2` before loaded into the capillary ELISA cartridge for the best ELISA results. NonOptimalCaptureAntibodyDiluent The CaptureAntibodyDiluent option should be kept as Model[Sample,"Simple Plex Reagent Diluent"] or a sample object with this model for the capture antibody samples `1` for optimized dilution and ELISA results. These capture antibody samples are used for ELISA assays of `2`. NonOptimalCaptureAntibodyDilution The specified CaptureAntibodyTargetCconcentration for the capture antibody samples `1` is higher than 50 Microgram/Milliliter. These capture antibody samples are used for ELISA assays of `2`. Please specify a lower concentration to achieve better ELISA test results. NonOptimalCaptureAntibodyPurificationColumn The specified CaptureAntibodyPurificationColumn should be a Zeba 40K MWCO spin column for the purification of capture antibody samples `1`, used for ELISA assay of the samples `2` for the best purification results. Please consider selecting `3`. NonOptimalDetectionAntibodyDiluent The DetectionAntibodyDiluent option should be kept as Model[Sample,"Simple Plex Reagent Diluent"] or a sample object with this model for the detection antibody samples `1` for optimized dilution and ELISA results. These detection antibody samples are used for ELISA assays of `2`. NonOptimalDetectionAntibodyDilution The specified DetectionAntibodyTargetConcentration for the detection antibody samples `1` is higher than 50 Microgram/Milliliter. These capture antibody samples are used for ELISA assays of `2`. Please specify a lower concentration to achieve better ELISA test results. NonOptimalDetectionAntibodyPurificationColumn The specified DetectionAntibodyPurificationColumn should be a Zeba 40K MWCO spin column for the purification of detection antibody samples `1`, used for ELISA assay of the samples `2` for the best purification results. Please consider selecting `3`. NonOptimalLoadingVolume The loading volume of samples `1` should be larger than 50 Microliter for optimized results. NonOptimalStandardCaptureAntibodyDiluent The StandardCaptureAntibodyDiluent option should be kept as Model[Sample,"Simple Plex Reagent Diluent"] or a sample object with this model for the standard capture antibody samples `1` for optimized dilution and ELISA results. These standard capture antibody samples are used for ELISA assays of the standard samples `2`. NonOptimalStandardCaptureAntibodyDilution The specified StandardCaptureAntibodyTargetConcentration for the standard capture antibody samples `1` is higher than 50 Microgram/Milliliter. These standard capture antibody samples are used for ELISA assays of the standard samples `2`. Please specify a lower concentration to achieve better ELISA test results. NonOptimalStandardCaptureAntibodyPurificationColumn The specified StandardCaptureAntibodyPurificationColumn should be a Zeba 40K MWCO spin column for the purification of standard capture antibody samples `1`, used for ELISA assay of the standard samples `2` for the best purification results. Please consider selecting `3`. NonOptimalStandardDetectionAntibodyDiluent The StandardDetectionAntibodyDiluent option should be kept as Model[Sample,"Simple Plex Reagent Diluent"] or a sample object with this model for the standard detection antibody samples `1` for optimized dilution and ELISA results. These standard detection antibody samples are used for ELISA assays of the standard samples `2`. NonOptimalStandardDetectionAntibodyDilution The specified StandardDetectionAntibodyTargetConcentration for the standard detection antibody samples `1` is higher than 50 Microgram/Milliliter. These standard detection antibody samples are used for ELISA assays of the standard samples `2`. Please specify a lower concentration to achieve better ELISA test results. NonOptimalStandardDetectionAntibodyPurificationColumn The specified StandardDetectionAntibodyPurificationColumn should be a Zeba 40K MWCO spin column for the purification of standard detection antibody samples `1`, used for ELISA assay of the standard samples `2` for the best purification results. Please consider selecting `3`. NonOptimalStandardDiluent The StandardDiluent specified for the standard samples `1` should be the preferred diluent `2` of the cartridge to achieve the best ELISA results. NonOptimalStandardDilutionCurve The total number of dilutions for standard samples `1` should be larger than 5 to generate a valid standard curve. NonOptimalStandardLoadingVolume The loading volume of standard samples `1` should be larger than 50 Microliter for optimized results. NonOptimalStandardResuspensionConcentration To achieve the best standard curve through sample manipulation of the standard samples `1`, it is recommended to set the resuspension concentrations to 10 times of the Upper Limit of Quantitation (`2`) of their analytes. NonOptimalUserSuppliedWesMolecularWeightRange The user-supplied MolecularWeightRange option, `1`, is not optimal for the average ExpectedMolecularWeight of the input antibodies, `2`. Please double check that the MolecularWeightRange option and antibody inputs are set as desired. NonOptimalWashBuffer The wash buffer should be Model[Sample,"Simple Plex Wash Buffer"] or an object with this model, as provided by the assay developer, for the best washing result. NonStandardSolvent The specified solvent(s) `1` are not one of ECL's currently supported solvents `2`. Note that this may result in a poor spectrum, and we recommend using a deuterated solvent. Nonsterile Some of the specified object (`1`) are non-sterile and will come into contact with your input samples (`2`), which are presently sterile. If you wish to avoid risking lost of sterilization, please consider using sterile samples in lieu of `2`. NonTabletSamples The field Tablet in the sample(s) `1` do(es) not contain tablets. Remove the sample(s) from the inputs or double check that the sample(s) do(es) contain tablets. NoReducingAgentIdentified A reducing agent was not identified in the composition of the reducing agent object specified for `1`. Please make sure that the object has valid composition or specify the approproate reducing agent volume. NoStatusChange The job(s) `1` already had Status `2`, and their status has not been changed. NotebooksIsNotSet PublicObjects cannot be set to False if Notebooks is not set and so public objects will be included in the search results. Please update your options and try again if this is not acceptable. NotEnoughBiotinReagentVolume The BiotinReagentVolume should provide excess amount of biotin reagent compared to CustomDetectionAntibody in the bioconjugation process to achieve the best conjugation efficiency for detection antibody samples `1` (used for ELISA experiment of `2`). NotEnoughDigoxigeninReagentVolume The DigoxigeninReagentVolume should provide excess amount of digoxigenin reagent compared to CustomCaptureAntibody in the bioconjugation process to achieve the best conjugation efficiency for capture antibody samples `1` (used for ELISA experiment of `2`). NotEnoughOptimalUsesLeftOnCartridge This protocol requires more injections than the cartridge `1` has left for optimal conditions (`2` injections). Please consider splitting this protocol or using another cartridge. NotEnoughOptimalUsesLeftOnCIEFCartridge This protocol requires more injections than the capillary Isoelectric Focusing cartridge `1` has left for optimal conditions (`2` injections). Please consider splitting this protocol or using another cartridge. NotEnoughStandardBiotinReagentVolume The StandardBiotinReagentVolume should provide excess amount of biotin reagent compared to StandardDetectionAntibody in the bioconjugation process to achieve the best conjugation efficiency for standard detection antibody samples `1` (used for ELISA experiment of `2`). NotEnoughStandardDigoxigeninReagentVolume The StandardDigoxigeninReagentVolume should provide excess amount of digoxigenin reagent compared to StandardCaptureAntibody in the bioconjugation process to achieve the best conjugation efficiency for standard capture antibody samples `1` (used for ELISA experiment of `2`). NotEqualBlankVolumesWarning The blank volume (`1`) is not equal to the volume of the sample (`2`). We recommend a blank volume that equals to the sample volume, or let the BlankVolumes resolve automatically. NotReplacingInsert The cartridge insert will not be replaced, even if required. This may affect the reliability of this cartridge the next time it is used. It is recommended to replace the insert if and when needed. Please consider setting ReplaceCartridgeInsert to True. NoTrigger Option Trigger was either not specified or set to None, so option TrackedFields will be ignored. NoVolumeIncrementsProvided The provided FulfillmentScale option was specified as `1`, but no VolumeIncrements were specified. Please provide VolumeIncrements if you would like to specify Fixed increments for fulfillment. ObjectDoesNotContain3DCurves The provided object/protocol does not contain data for 3D melting/cooling curves. Please provide only Object[Protocol, UVMelting] and Object[Data, MeltingCurve] that contain 3D data. Attempting to use 2D curves for this analysis. OnBoardMixingVolume Onboard mixing requires a minimum volume in each vial in order to run reliably. As a result, an extra `1` of the prepared sample mix will be required. OnlyOneAgaroseLoadingDye The following input samples, `1`, have only one associated LoadingDye specified. Sizing is most accurate when each SampleIn has two associated LoadingDyes whose oligomer lengths flank the target strand of interest. Please consider specifying an additional LoadingDye for this input, or letting the LoadingDye option automatically resolve. OnlyOneCalibrantWillBeRan If the MassAnalyzer is not TripeQuandrupole, only one calibrant will be ran during the experiment: `1`. The SecondCalibrant (`2`) will not be ran. OptionPattern The given option value `1` does not match the pattern `2` for the option `3` in the function `4`. Defaulting the value to `5`. OptionUnavailable Option `1` cannot be used with resolved PeakType `2`, and will be ignored in analysis. OsmolalityInoculationPaperHighViscosity In ExperimentMeasureOsmolality, the samples, `1`, are specified/resolved to use viscous loading techniques, and it is specified to use inoculation paper. If your sample is viscous, consider not using sample paper as the sample may not be effectively absorbed by the paper and more even distribution may be achieved by application directly to the sample holder, producing a more reliable measurement. OsmolalityLowSampleVolume In ExperimentMeasureOsmolality, the sample volumes, `1`, are lower than the minimum recommended volume for a reliable reading with `2`, of `3`. Consider specifying a larger volume of sample, or take this into consideration when interpreting the result. OsmolalityNoInoculationPaper In ExperimentMeasureOsmolality, the samples, `1`, are specified/resolved not to use viscous loading techniques, and it is specified not to use inoculation paper. If your sample is not viscous, consider using inoculation paper as it helps to ensure an even distribution of sample in the sample holder, producing a more reliable measurement. OsmolalitySampleCarryOver In ExperimentMeasureOsmolality, the samples, `1`, are not viscous, however ViscousLoading is set to True. Viscous loading techniques have a higher risk of carry-over error, and therefore should only be used when necessary. Please consider setting ViscousLoading to False if not required. OsmolalityShortEquilibrationTime In ExperimentMeasureOsmolality, the equilibration times for samples `1` on instrument `2` are timed manually by the operator and so short equilibration times, `3`, will have higher uncertainty. Please consider specifying a longer equilibration time if precise timing of the equilibration period is important. OsmolalityUnknownViscosity In ExperimentMeasureOsmolality, the viscosity of the samples `1` could not be found to determine the appropriate loading procedure for the instrument. The standard loading procedure for low viscosity samples is assumed - if your sample is not viscous, set ViscousLoading to False to silence this warning. If your sample is viscous, set ViscousLoading to True to avoid issues with loading. If you are unsure whether your sample is highly viscous, consider running ExperimentMeasureViscosity. OsmolalityViscousTransferMinimumVolume In ExperimentMeasureOsmolality, the samples, `1`, are specified/resolved to use viscous loading techniques which utilize a positive displacement pipette with a minimum measurable volume of 10 uL. The volumes `2` will not be measured accurately. If possible, consider specifying a sample volume of 10 uL or more. OutdatedTemplateOptions The template protocol contains options which don't match their pattern. These options cannot be used and will be set to their default values. If you don't wish to use the defaults, please directly specify new values for these options. Check: `1` OutstandingResources The samples `1` set for Disposal have Outstanding resource requests; these samples will not be discarded until these resources are no longer outstanding. OveraspiratedTransfer The source(s), `1`, have the following amounts specified to be transferred, `2`, at manipulation indices `4`. However, these source(s) will only have `3` in their containers at the time that the transfer is to be performed. Please specify a lower mass/volume/count or specify All to transfer all of the contents of the sample. OverwriteLadderStorageCondition A Ladder is not used in this experiment. There LadderStorageCondition is overwritten and set to Null. OverwritingGradient The inherited gradient will be overwritten based on setting of other options: `1`. Please review and ensure that it meets desired specifications. OverwritingMassAcquisitionMethod The following acquisition methods specified in the following options `1` will be overwritten with new methods because of specified acquisition changes. OverwritingSamplelessChannelOptions One or more `1` related options are specified while there's no `1`. These options are overwritten and set to Null. OverwritingSeparationMethod The specified SeparationMethod will be overwritten based on setting of other options: `1`. Please review and ensure that it meets desired specifications. ParallelProjectionInvalid Parallel projection of a single contour is not supported. In the current plot, WaterfallProjection has instead been set to Perspective. If parallel projection is desired, please try adding more contours. ParameterMismatch Supplied value `1` for option `2` does not match value `3` found in input `4`. Please verify that `1` is correct, or use Automatic to default to input value. ParentPeakIndices Indices `1` in option ParentPeaks are outside of the range 1 to `2`, the number of identified peaks, and will be defaulted to the largest peak (index `3`). ParentPeaksMismatch Parent peaks contain labels `1` which do not correspond to any labels or assignments. Please ensure that all entries in parent peaks have a corresponding label or assignment. ParentPeaksPadded `1` parent peaks were specified, but `2` peaks were identified. Extra parents will be ignored, and unspecified parents will default to the largest peak. PeakGroupMultiplicityMismatch The number of peak multiplicities (`1`) does not match the resolved number of peak groups (`2`). Excess multiplicities will be ignored, and missing specifications will be defaulted. PeakLabelsMismatch Number of Peak labels does not match number of peaks. Excess labels will be ignored. pHCalibrationTarget The specified SecondarypHCalibrationBufferTarget does not match the pH from the SecondarypHCalibrationBuffer packet. Please consider letting the SecondarypHCalibrationBufferTarget set automatically. PlateReaderStowaways There are additional samples in the plate being experimented upon. As a result they will also be subjected to the heating and/or mixing done in the plate reader chamber. If you don't wish these samples to be acted on, please set Aliquot->True. PooledConsolidateAliquotsNotSupported The ConsolidateAliquots option has been set to True when pooling multiple samples together. Consolidating aliquots is not currently supported when pooling multiple samples together. Option resolution will continue as if ConsolidateAliquots is set to False. PooledVolumeBelowMinThreshold For the input sample(s) `1`, the total volume of the pooled sample(s) and buffer(s), `2`, is below the minimum volume at which the chosen AliquotContainer, `3`, can be reliably read (`4`). If you wish to acquire data within the recommended working range, specify a smaller AliquotContainer or leave this option Automatic. Please refer to the documentation for a list of available cuvettes and their working volumes. PotentialsPrecision At least one member in the provided `1` has higher precision than the instrument specifications. They have been rounded from `2` to `3`, with the supported precision of 1 Millivolt. PotentialSweepRatePrecision At least one member in the provided PotentialSweepRate has higher precision than the instrument specifications. They have been rounded from `1` to `2`, with the supported precision of 10 Millivolt/Second. PowderXRDHighVolume The volume of input samples `1` (or their resolved AssayVolume) (`2`) is above 100 microliters. If using a sample that is a solid suspended in a solvent, we recommend using volumes at or below 100 microliters to ensure the suspension is adequately concentrated such that sufficient solid may be transferred to the XRD plate. PredictedValueOutOfDomain Warning, values predicted by the standard curve function are outside the range of the standard curve data. Extrapolation is being used. PreferredIlluminationIncompatible The sample(s) `1`, are in containers whose PreferredIllumination is not possible on their specified imaging instrument(s), `2`. A compatible illumination direction has been selected for these samples. PreMadeMasterMixWithMakeOwnOptions Options for both premade MasterMix and modular MasterMix are specified for `1`. These options are exclusive to each other and only premade master mix options will take effect. If you dont wish to use a PremadeMasterMix, make sure to set this option to False. PRENullPrimaryWashingSolutionWarning The PrimaryWashingSolution is set to Null for the following input target reference electrode models `1`. Please consider using a washing solution to wash the reference electrode before the priming step. PreparedPlateFalse The samples `1` are already in a tensiometer assay plate `2` and PreparedPlate is set to `3`. If you would like to measure the plate without any extra sample manipulation, set PreparedPlate to True. PREReferenceCouplingSampleSwitchingWarning The following input target reference electrode models `1` have a different reference coupling sample from their corresponding source reference electrodes. Please consider use a different source reference electrode model or object. PRESourceReferenceElectrodeWorkingPartRustWarning The following input source reference electrode objects `1` have rust observed on its working part (the part that directly contacts experiment solution) and should be discarded. Please consider use another reference electrode for the preparation and use the DiscardSamples function to discard the current reference electrode. PressureVentingTriggersPrecision The PressureVentingTriggers values were provided at higher precision than can be executed in this experiment. The values `1` have been rounded to `2`. PricingObjectsArePrivate Full pricing information cannot be displayed because the following object(s) are no longer accessible by members of the current financing team as they may have changed ownership: `1`. Please note that these items may still result in usage charges. PrimaryDataInvalid One or more of the input data objects do not contain 2-D or 3-D data within the PrimaryData field (`1`). Please check that PrimaryData refers to a field containing 2-D data or 3-D data suitable for plotting. Defaulting to the first available field containing valid data. PrimersNotFound No PCR primers could be resolved for the provided input. Please provide a TerminationSequence, or change TrimmingMethod from PrimerTerminus to another option. PrimitivesWithIncompatibleKeys The primitive at index `1` has a key(s) that is not supported by ExperimentAcousticLiquidHandling. Keys that are not Source/Sources/Destination/Destinations/Amount/Amounts/InWellSeparation will be ignored. ProtocolAlreadySet The option Protocol is redundant when the input is of type Object[Protocol]. Protocol will be ignored. PulseSequenceSpecified A custom pulse sequence is specified for sample(s) `1`. This will override whatever the ExperimentType option is set to. Note that this option should only be set by pro users and that the user is liable for any damage this custom pulse sequence may inflict on the probe or instrument. QuantificationCycleNotFound No quantification cycle within the amplification range is found for template sample(s) `1` and is being set to 0 Cycle. If a quantification cycle is expected, please check and adjust the values of BaselineDomain and/or Threshold. RamanSampleWithoutBlank Samples `1` do not have specified Blank. If you wish to subtract out background signal arising from the plate or any solvent or solid matrix, specify an appropriate Object or Window for the Blank option. RamanSamplingCoordinatesPrecision The provided SamplingCoordinates have higher precision than the instrument specifications. They have been rounded to the nearest Micrometer. ReactionTypeNull Input Object `1` should not have ReactionType option set to Null. Changing ReactionType option to `2`. ReactionTypeWarning Object `1` may not have bonds and option `2` is selected. You may want to set ReactionType option to Automatic. RecommendedInstrument The specified instrument `1` is not recommended for the input sample analytes. Consider using `2` instead. RecoupSuspensionSolution For the following sample(s): `1`, Recouping is requested along with a SuspensionSolution. This will add the SuspensionSolution to the input sample. ReflectionOnly Transmission sample module was requested. Currently, we only offer Reflection sample module for the sample loading and measurement. The experiment function will proceed with this. RelativeMigrationDataUnavailable No RelativeMigrationData was found for inputs `1`. Please run AnalyzePeaks on these inputs and use the ParentPeaks option to specify the internal standard peak which should be used as a reference for RMT. ReloadSLL The version of the `1` manifest loaded in the Mathematica Kernel is not the same as the version of the `1` manifest present in the local `1` folder. Please reload SLL to avoid creating conflicts between the two versions. RemovedExtraGradientEntries A specified gradient in `1` after rounding to acceptable time had duplicate entries. The duplicates were removed. RepeatedDetectors The specified Detector option `1` has repeated entries. The repeat entries have been removed. RepeatedPlateReaderSamples Note that in order to generate readings of samples repeated multiple times in the input, samples will be aliquoted into a new plate. ReplicateAliquotsRequired Note that in order to generate replicate readings samples will be aliquoted into a new plate. ReplicateChipSpace When concentration is being quantified, 3 replicates are recommended, however there is insufficient space in the chip to do this. NumberOfReplicates will be set to `1`, the largest possible value, but you may wish to consider splitting this into multiple experiments. RequiredIntermediateAliquot `1` resolveSamplePrepOptionsNoCache resolveSamplePrepOptions needs to be passed cache in order to function correctly. Please pass down packets for all of the containers and samples that you are using. ReverseComplexCoupling Automatic calculation of J-coupling constants is not supported for complex multiple splittings. `1` peaks were expected for the specified multiplicity `2`, but only `3` were identified in the splitting group. RotorDefaultStorageConditionChilledRotorConflict For samples `1`, ChilledRotor option set to True, it is desired to pick Rotor that are stored in Refrigerator. Please adjust either options. RoundedCorrectionCurve CorrectionCurve values cannot exceed a precision of 0.1 Microliter. The specified values `1` will be rounded to the nearest 0.1 Microliter. RoundedTransferAmount The give amounts to be transferred, `1`, at manipulation indices, `3`, are going to be rounded to, `2`, due to the Resolution of the transfer Instrument/Balance that are going to be used. If the given precision is still desired, please transfer a smaller amount or use a different Instrument/Balance to perform the transfer. SameCalibrantsForLCMS The specified carlibrants `1` and `2` are same, only one calibrant will be ran in experiment. SampleAndContainerSpecified The following sample(s) were specified along with the containers holding these sample(s), `1`. If this was intended, this message can be ignored. If you do not wish to operate on these samples twice, please remove them or their containers from your input list. SampleMassMayBeInaccurate For the following input container(s), `1`, CalibrateContainer is specified as True, while it contains a sample with a recorded Mass that may be outdated since other protocols have manipulated this sample after the latest weight measurement. Please consider leaving CalibrateContainer Automatic in order to record the accurate mass of the the sample before calibrating the container. SampleMayBeTransferred Although a volume was specified for these models, `1`, the density is not known. If the sample arrives in a container that is not parameterized for liquid level detection, your sample will be transferred to another container upon arrival. If this is not desired, please populate the Density of the models. SampleMustBeMoved The experiment requests sample `1` to be put in container model(s) `2`. Because `1` is not currently in a container with this model(s), this protocol will transfer the designated amount to a new container with this model(s) SamplesAlreadyDiscarded Unfortunately, the samples `1` have already been permanently thrown out and can no longer be undiscarded. These samples have been excluded from these updates. SamplesInTransit The experiment requests samples that currently have the status ShippingToECL (`1`). The protocol will not start in the lab until these samples arrive. SamplesMarkedForDisposal The following object(s) specified are flagged for disposal: `1`. If you don't wish to dispose of these samples, please run CancelDiscardSamples on them. SamplesNotOwned The following object(s) specified are not part of a Notebook financed by one of the current financing teams: `1`. Please check the Notebook field of these objects and the Financers field of that Notebook; if they are public, please purchase these items before directly requesting them in an Experiment. SamplesOutOfMassDetection The molecular weights of `1` fall outside the specified mass range. In order to capture data about these samples, you may wish to update your mass ranges. SamplesOutOfStock The following input model(s) `1` have no currently available non-expired samples in the ECL, but they can be ordered and we estimate they will arrive in approximately `2` after the item is ordered. SamplesOutStorageConditionMismatch The SamplesOutStorageCondition is set to `1` but ContainerOut is specified `2`. If you would like to store your samples, please specify a SamplesOutStorageCondition. SampleStowaways The input samples to this experiment reside in container(s) `1`, however these containers contain additional samples (`2`) which weren't provided as input. In order to avoid modifying these additional samples, the input samples will be transferred into a new container before beginning the experiment. If this is undesired please re-run the experiment function using the container you wish to run the experiment on as the input. SamplesWithDeprecatedModels The following sample object(s) specified have models that are deprecated: `1`. Please check the Deprecated field for the models in question, or provide samples with alternative models. ScanTimeAdjusted The scan time specified in the following options do not match the acceptable values and will have to be rounded to proceed: `1`. If you don't wish rounding to occur automatically, please supply a value from the following list: `2`. ScatterPlotOverride When a secondary variable is given with AlphaScreen data in PlotAlphaScreen, only ScatterPlot can be plotted. The specified PlotType `1` will be ignored. ScheduleDateCannotBeDetermined A schedule for the following target - systems protocol model pairs could not be determined (`1`). This input has been excluded. Please confirm that this systems protocol model is run on a schedule for this target. ScheduleSmallGap The consolidated StorageSchedules of objects `1`, contains consecutive entries with time gaps of less than 12 hour. Timing of the storage update is difficult to be guaranteed in this case. We recommend choose a larger gap using Date option or use ClearSampleStorageSchedule to clear StorageSchedule. SelectComponentsNotSpecified MinComponentRadius, MaxComponentRadius, AreaThreshold, or IntensityThreshold have been specified without using SelectComponents in the list of ImageSegmentation for Input `1` image index `2`. Please add SelectComponents[] to the end of ImageSegmentation to take these criteria into effect. SimilarMolecules The following molecules, `1`, have a same molecular identifier as the molecules, `2`, you are trying to upload. Please use this existing molecular model if suitable. SinglePlateRequired Note that all samples will be aliquoted into a single plate supported by the plate reader. SomeWavelengthsOutOfRange At least one of the specified wavelength values is not in the wavelength range that the data has been acquired with. These wavelength values are being ignored in the analysis. The available wavelength range in this data set is from `1` nm to `2` nm. SpacingToleranceExceeded SplittingSpacingTolerance exceeded while assigning multiplicity `1` to peaks at `2`. Please verify that the multiplicity assignment is correct, or increase SplittingSpacingTolerance. SpanWavelengthOrder The span wavelength (`1`) is specified from high-wavelength to low-wavelength. The instrument will automatically adjust and scan the samples from low-wavelength to high-wavelength. SplittingIgnoreUnits Input data was provided with x-units of `1` instead of PPM. Units will be ignored in peak-splitting analysis. StandardCaptureAntibodyDilutionRecommended StandardCaptureAntibodyDilution is recommended (not Null) for standard capture antibody samples `1` (used for ELISA assay for standard samples `2`), because resupension and/or conjugation of the antibody is performed in this protocol and the concentration of the antibody may be too high for capillary ELISA experiment. StandardConcentrationIncompatibleUnits Two or more of the units in the option StandardConcentrations `1` are incompatible. Please specify compatible units for all standards. The StandardConcentration option will be ignored, and standard concentrations will be computed from the Composition fields of Standards in the input protocol. StandardDetectionAntibodyDilutionRecommended StandardDetectionAntibodyDilution is recommended (not Null) for standard detection antibody samples `1` (used for ELISA assay for standard samples `2`), because resupension and/or conjugation of the antibody is performed in this protocol and the concentration of the antibody may be too high for capillary ELISA experiment. StandardFieldOverride `1` has been doubly specified in both inputs and options. Values specified through options will take precedence. StandardPrimerPairNotAmongInputs The specified standard primer pair(s) `1` do not appear in the set of input primer pairs and thus may not be useful for the quantification of experimental results. SterileConflict Sterile was set as False but some samples are marked as sterile. The experiment will proceed with a non-sterile instrument. If this is not desired, please adjust the sterile option. SterileContainerRecommended The sample `1` was requested to be sterile filtered. The provided FiltrateContainerOut option `2` is not sterile, but will be used. SuperloopInjectionVolumesRounded Volumes must be specified in increments of 0.01 Milliliter if InjectionType -> Superloop. Specified volumes `1` have been rounded to the nearest 0.01 Milliliter quantity. SwitchedKineticsModel The kinetic parameters of the polymer `1` is not available. The parameters of DNA will be used instead. SwitchedPlotType The PlotType `1` is not available, and will be switched to the one that is available in the analysis packet. SystemMessages `1` TabletCrusherRequired The manipulation(s) at indices `2` specify that a mass of the source sample, `1`, should be transferred to the destination. However, the source sample has the SampleHandling category of Itemized. A pill crusher may be used in order to achieve the requested mass. If this is not intended, please specify an integer number of pills instead of a mass to be transferred. TabletWeightKnown For sample(s) `1`, the option `2` is specified although the TabletWeight is known. As a result the tablets will be re-parameterized and the TabletWeight overwritten with the newly recorded value. Consider leaving `2` blank if you would like to keep the currently existing TabletWeight. TargetConcentrationNotUsed The TargetConcentration option has been specified for the following sample(s): `1` for which Amount has been specified as a Count: `2`. TargetConcentration is not used if specified for counted items. TargetConcentrationPrecision At least one member of the provided `1` has higher precision than the experiment specifications. It has been rounded from `2` to `3`. TemplateOptionUnused The provided Template option `1` is not the same object as the template input `2`. The input `2` will be used as the template. TooLateToCancelMaintenanceStorageUpdate StorageSchedules for samples `1` contain Object[Maintenance,StorageUpdate] that has already started. We cannot clean this entry of StorageSchedule but have cleaned the rest. tooManyMultpleReactionMonitoringAssays For samples `1` in ESI-QQQ, the input MultipleReactionMonitoringAssays has a length of `2`, this may influence the quality of the data. Please consider separating this experiment into several separate MultipleReactionMonitoring experiments for higher quality data. TotalAliquotVolumeTooLarge The following sample(s) `1` have a volume requested `2` that exceed the volume of these sample(s). The experiment will still attempt to aliquot these amounts with what is currently available; please change the Volume option if this is not desired. TotalColumnVolumeDefault As neither the BedVolume nor the ResinLoadingAmount has been specified, the BedVolume has been set to the TotalColumnVolume. The ResinLoadingAmount will be backcalculated from the BedVolume and the value of the SpecificVolume for the set resin and storage buffers, `1`. TotalProteinQuantificationDuplicateProteinStandards The specified ProteinStandards, `1`, has duplicate entries. Duplicate members of ProteinStandards will result in more absorbance or fluorescence data points in the StandardCurve for the associated protein concentration than the other protein concentrations. Please ensure that this duplicate entry is desired. TotalProteinQuantificationLoadingVolumeLow The LoadingVolume has been automatically set to `1`, which is lower than `2`, the ideal volume for the DetectionMode (`3`) and AssayType (`4`) options. This is because the QuantificationReagentVolume was set to `5`, and the sum of the LoadingVolume and QuantificationReagentVolume cannot be larger than 300 uL. Please consider setting the QuantificationReagentVolume lower, or letting the option resolve automatically. TotalProteinQuantificationMultipleProteinStandardIdentityModels The specified ProteinStandards, `1`, do not all have the same Model[Molecule,Protein]s in their Composition fields. The lists of Model[Molecule,Protein]s present in each ProteinStandard are `2`. The StandardCurve generated from these ProteinStandards may not be able to accurately quantify the concentration of proteins in the input samples. Please ensure that the ProteinStandards option has been specified correctly, or consider letting the option resolve automatically. TotalProteinQuantificationQuantificationReagentVolumeLow The QuantificationReagentVolume has been automatically set to `1`, which is lower than `2`, the ideal volume for the Detection Mode (`3`) and AssayType (`4`). This is because the LoadingVolume was set to `5`, and the sum of the LoadingVolume and QuantificationReagentVolume cannot be larger than 300 uL. Please consider setting the LoadingVolume lower, or letting both options resolve automatically. TotalProteinQuantificationReagentNotOptimal The specified QuantificationReagent, `1`,is not the default QuantificationReagent Model for the AssayType, `2`. Please ensure that this is desired. If it is not, consider letting the QuantificationReagent option automatically resolve. TotalProteinQuantificationTotalVolumeLow The sum of the LoadingVolume, `1`, and QuantificationReagentVolume, `2`, is less than 60 uL. Absorbance or Fluorescence values from such a low volume might not be accurate. Please consider increasing these volumes, or letting these options automatically resolve. tracePlumbingRecursionLimit tracePlumbing has hit the recursion limit of `1`. The list of objects may be incomplete. Please increase this limit or double check the plumbing network. TransactionIsNotSet Can not close unopened transaction. No open transactions TransferRequired The container of the input sample(s) `1` does not fit on the cuvette block of the spectrophotometer `2`, therefore the sample is going to be transferred into `3`. If you wish to avoid this transfer, please move the sample(s) to a cuvette prior to the experiment. Please refer to the documentation for a list of compatible containers. TransferTypeDefaulted The manipulation `1` was specified with TransferType `2`. However, this transfer type is inconsistent with the amount(s) of the manipulation. This manipulation will be performed via transfer type `3` TrimDirectionIncompatible The requested Trimming direction `1` is incompatible with TrimmingMethod `2`, and results will be calculated as if Trimming->`3`. Please change the option Trimming to one of `4`, or choose a different TrimmingMethod. TrimImpossible No satisfactory trimming could be found for the inputs at indices `1`. Please change the direction of the Trimming option, increase trimming thresholds, or choose a different TrimmingMethod. TrimmingOptionsIgnored The trimming options `1` were specified but Trimming was either not set or set to None. Trimming options will be ignored. Please change Trimming to StartEndBoth, or remove these options. UltracentrifugeChilledRotorHighTempConflict For samples `1`, ChilledRotor option set to True, it is desired to set the Temperature option to be between 4 Celsius and Ambient. Please adjust either options. UltracentrifugeChilledRotorLowTempConflict For samples `1`, ChilledRotor option set to False, it is desired to set the Temperature option to be Ambient to 40 Celsius. Please adjust either options. UnableToConvertConcentration The concentration of the TargetMolecules `1` in the samples can not be converted to Molar. If you would like to convert the these concentrations, please make sure the MolecularWeight of `1` is uploaded. UnableToExpandInputs None of the provided inputs in the IndexMatching block `1` have been expanded so they will be returned unchanged. UnableToExpandOptions The provided options cannot be expanded and will be returned unchanged. If you wish to expand these options at least one must be expanded so that the expansion length can be determined. The unexpanded options are: `1`. UnableToFindExactSLLCommit We were unable to find a built commit for `1`. Instead, we will use the latest commit with an analyzed codebase, `2`. Please run ReloadPackage[myPackage] or ReloadFile[myPackage, myFile] to re-analyze any packages that might have changed between these commits, after you've loaded in the pre-cached analyzed package. UnableToFindPreviousAnalysis We were unable to find a pre-cached version of SLL for this branch. We will manually analyze the codebase from scratch (this will take a while). To speed up the process, make sure that you have a pull request open for your branch so that Neutrino automatically builds analysis information for each commit. UncertainMeltingOnset The onset value determined for `1` might not be accurate. This is likely related to the shape of the curve which can be deviated from sigmoid-like. UndatedEnvironmentalChamberStorageEnqueued Samples `1` are enqueued to be placed into an environmental chamber but Date is not specified. The sample will be put into the environmental chamber at the next daily lab-wide storage update. If you prefer this to be a dedicated, timed operation, please specify the Date option. Use ClearSampleStorageSchedule to clear all entries of StorageSchedule. UndefinedFlowRate The flow rate is not specified. Plot will be generated without the transform. UndetectableFluorescentDetectionLabels Mode `1` will not be able to detect the following fluorescent molecules specified in DetectionLabels option at index `2`: `3`. Please select the following Mode to allow detection of fluorescent DetectionLabels: `4`. UndetectableNonFluorescentDetectionLabels Mode `1` will not be able to detect emitted fluorescence from the following molecules specified in DetectionLabels option at index `2`: `3`. Please select the following Mode to allow detection of non-fluorescent DetectionLabels: `4`. UndeterminedSpecificVolume The set resin and storage buffers, `1`, do not have a determined specific volume. Therefore, the conversion coefficient between BedVolume and ResinLoadingAmount will be determined during the protocol. UnknownAmount The following sample(s) do not have a known current volume or mass: `1`. Additional checks to validate the requested aliquot amounts will not be performed for these samples. UnknownAnalytes Unable to identify protein or nucleotide analytes. Other types of analytes are not recommended for this assay. Automatic options will default to oligomer analyte settings. UnknownComposition No chemical formula or structure is defined for monomers `1` of type `2`. Isotopic variation will be ignored for these monomers and their average molecular weight will be used instead. UnknownMasterMix The specified master mix model, `1`, has not been evaluated for use in ExperimentqPCR and has unknown concentration, dye content, and reverse transcriptase content. Unless specified otherwise, the master mix will be used at 2x concentration and will be assumed to contain `2`. Please refer to the ExperimentqPCR documentation for a list of known master mixes. UnknownMolecularWeight The molecular weight of the analytes in the sample(s), `1`, is unknown. As a result a default mass range will be chosen, and the analyte(s) of interest may not be detected if it falls outside of the range. If you have a sense of the expected weight, please consider providing a value for the option MassDetection. UnknownMonomer The phosphoramidite to use for the monomers `1` which are in the strands `2` could not be determined. Please specify the phosphoramidite to use for the monomer in the Phosphoramidites option, or synthesize oligomers that do not use this monomer. UnknownOption The following specified option(s) are not options for `2`: `1`. UnknownProductStoichiometry The following samples do not have defined ProductStoichiometricCoefficient: `1`. The stoichiometric coefficient will default to 1 for the corresponding product. If this is not accurate please update ProductStoichiometricCoefficient. UnknownReactantsStoichiometry The following samples do not have defined ReactantsStoichiometricCoefficients: `1`. The stoichiometric coefficient will default to 1 for each reactant. If this is not accurate please update ReactantsStoichiometricCoefficients. UnknownUnits Unknown standard units for `1`. UnmatchedAssignment The assignment(s) `1` in option SplittingAssignments could not be matched to any resolved peak splitting domains, and will be ignored. UnnecessaryBlankTransfer Although they are already in compatible containers, blanks wil be transferred from their current container into new containers. If you don't wish for these samples to be transferred set BlankVolumes->Null for `1` UnneededBLIProbeRegeneration Only one measurement step is performed, so there is no need to regenerate the probe. UnneededSyringeComponent The specified `1` is not required, as none of the values in option(s) `3` specified are `2`. Please ensure option `3` has been specified correctly for each sample. UnrecognizedOption Option(s) `1` are not valid options for this command and have been ignored. Please double-check all your options are spelled correctly. UnrecognizedPeaksField Unable to interpret given ReferenceField. Please check the ReferenceField is correct for the input data type. UnresolvableEmissionWavelength The EmissionWavelength option at index `1` is set to `2`, which is the longest wavelength supported by the instrument because DetectionLabels option is Null. Please specify the EmissionWavelength option if a different wavelength is preferred. UnresolvableExcitationWavelength The ExcitationWavelength option at index `1` is set to `2`, which is the longest wavelength supported by the instrument because DetectionLabels option is Null. Please specify the ExcitationWavelength option if a different wavelength is preferred. UnresolvableFlowCytometryDetector The provided DetectionLabel (`1`) did not match any of the detectors for this instrument. UnresolvedPlotOptions Resolved option(s) `1` are equal to Automatic. Please ensure that these options have resolved correctly. UnsealedShelfLifeLonger The provided UnsealedShelfLife value(s) `1` are longer than the provided ShelfLife value(s) `2`. UnspecifiedUnits Data supplied in `1` has unspecified units, which are assumed to be compatible with units `2` in `3`. Please verify that the data supplied in `1` is correct. UnsupportedMethod Unable to use Base method on degenerate sequences due to vagueness of pairing rules on degenerate sequences. Will default to Motif method. UnsupportedOption Option(s) `1` are not supported by the Command Builder and have been ignored. UnsupportedTrimmingMethod The requested TrimmingMethod (`1`) is not supported if UnassignedBases->False. Please set UnassignableBases->True, or choose a different trimming method. UnsupportedUnits Unsupported units `1` detected in input. Units will be ignored in analysis. Please ensure that input xyData is either unitless, or has units of ArbitraryUnit. UnusedBLIEquilibrateOptions The following options are only used if Equilibrate -> True: `1`. Change the value of Equilibrate to True or Automatic if and equilibration step is required. UnusedBLINeutralizeOptions The following options are only used when RegenerationType includes Neutralize: `1`. UnusedBLIProbeEquilibrationOptions The following options are only used if ProbeRackEquilibration -> True: `1`. Change the value of ProbeRackEquilibration to True or reset these options to Automatic or Null. UnusedBLIRegenerationOptions The following options are only used when RegenerationType includes Regenerate: `1`. UnusedBLIStandard The value of Standard has been overridden by QuantitationStandard. If a more complicated assay is desired, consider using the AssaySequencePrimitives or ExpandedAssaySequencePrimitives options. UnusedBLIWashOptions The following options are only used when RegenerationType includes Wash: `1`. UnusedComponentProperty The Property is unused for the primitive `1` of input `2`. If Criteria field is specified for the SelectComponents primitive, the Property field will be set to Null. UnusedCutoffFrequency The provided CutoffFrequency for dataset `1` is not used. The method `2` does not take CutoffFrequency information. Consider using Radius for manual control over the smoothing strength UnusedDataSet The DataSet is set to Null if the temperature and response datasets are not Null or if the input pattern is {{object1,object2,..}..}. Please specify TemperatureDataSet and ResponseDataSet if you wish to set the field that the data points should be taken from. UnusedDataSetTransformation The DataSetTransformationFunction is set to Null if the temperature and response datasets are not Null or if the input pattern is {{object1,object2,..}..}. Please specify TemperatureDataSet and ResponseDataSet and their TransofrmationFunction if you wish to set the field that the data points should be taken from. UnusedLiquidHandler The LiquidHandler option was specified, but the manipulation scale has defaulted to MacroLiquidHandling; the provided liquid handler option will not be used. UnusedOptimizationParameters For the following samples, `1` option(s) will be ignored because OptimizeFluorescenceLaserPower is False:`2`. Please set OptimizeFluorescenceLaserPower to True if laser power optimization is desired. UnusedOptionValuesBLIPrimitiveInput The following options were specified, but are overridden by the user input AssaySequencePrimitives: `1`. UnusedProperty The Property, OrderingCount, and OrderingFunction is unused for the primitive `1` of input `2`. If Criteria field is specified for the SelectComponents primitive, the Property, OrderingCount, and OrderingFunction fields will be set to Null. UnusedRadius The provided radius for dataset `1` is not used. The method `2` does not take Radius information. Consider using CutoffFrequency for manual control over the smoothing strength. UnusedResponseTransformation The ResponseTransformationFunction is used if TemperatureDataSet and ResponseDataSet are not Null. Please specify DataSetTransformationFunction if you wish to apply a transformation to the DataSet or specify TemperatureDataSet and ResponseDataSet if you wish to set the field that the data points should be taken from. UnusedSquarePosition Method for input `1` is set to Manual or Hybrid and therefore HemocytometerSquarePosition is set to All. Please use Method->Automatic if automatic detection for a specific square is intended. UnusedTemperatureTransformation The TemperatureTransformationFunction is used if TemperatureDataSet and ResponseDataSet are not Null. Please specify DataSetTransformationFunction if you wish to apply a transformation to the DataSet or specify TemperatureDataSet and ResponseDataSet if you wish to set the field that the data points should be taken from. UnusedTemplateOptions The template protocol contains options which don't currently apply to this experiment and so these options cannot be used. Please check the options of `1` and compare with the resolved options of the template protocol. Check: `2` UnusedThresholds The DetectionThresholds option `1` cannot be used because input protocol has UnstainedSampleData which is being used as a global negative. The DetectionThresholds option will be ignored. UploadQuizQuestionSynonymMissing In UploadQuizQuestion, for questions `1`, the Synonym options provided do not contain the name provided by the Name option. This name will be prepended to the synonyms. UREMismatchingSolventTypeWarning The ReferenceSolution has a mismatching solvent type with the RecommendedSolventType, which means the ReferenceSolution has water has its solvent while the RecommendedSolventType is Organic, or the ReferenceSolution has an orgniac solvent while the RecommendedSolventType is Aqueous. Please consider matching the ReferenceSolution solvent and the RecommendedSolventType. UVVisOptionsNotApplicable The following options `1` are not available for the UVVis detector on the instrument best suited to meet all options. These options will be ignored. VariableColumnTypes The specified Columns `1` have a SeparationMode of `2`. When using Column SeparationMode to resolve automatic options, only the SeparationMode for the first column will be considered. ViewPointUnitMismatch A dimension of the specified ViewPoint (`1`) has units that are incompatible with the same dimension of the input data (`2`). Please check that the ViewPoint units are consistent with the input data, or specify a unitless numerical value. Defaulting to the provided numerical value (`3`). ViscosityCustomMeasurementTableNotProvided In ExperimentMeasureViscosity,the MeasurementMethod is specified as Custom, but no custom measurement parameters are provided in the MeasurementMethodTable. Please input the desired measurement parameters or choose another MeasurementMethod. ViscosityFlowRateSweepInsufficientInput In ExperimentMeasureViscosity, only one FlowRate, `1`, is specified for sample `2`, but the MeasurementMethod was specified as FlowRateSweep. In order to conduct a FlowRateSweep, please provide at least 2 FlowRates or set FlowRate to Automatic. ViscosityNotEnoughSampleInjected In ExperimentMeasureViscosity, the calculated total volume required for taking all measurements for `1` exceeds the resolved InjectionVolume, `2`, and the instrument may not be able to complete all measurement steps. Please specify a larger InjectionVolume or set RemeasurementAllowed to True so that the sample may be recycled and pumped back into the meausrement chip for additional measurements. ViscosityPressureMeasurementTimeConflict In ExperimentMeasureViscosity, the provided MeasurementMethodTable `1` for sample `2` contains one or more measurement steps where RelativePressure and EquilibrationTime and/or MeasurementTime are specified. EquilibrationTime and MeasurementTime are dependent on FlowRate, which will be determined by the instrument at experiment time. In order to allow the instrument determine the optimal EquilibrationTime and MeasurementTime for a given RelativePressure, please set these values to Automatic or Null. ViscosityPrimingFalse In ExperimentMeasureViscosity, Priming is False for sample(s) `1`. Priming is highly recommended to initially wet the measurement chip and before measuring at each new MeasurementTemperature. VolumeOptionUnused The Volume option was set to the specific volume `1`, but the provided formula has a total/fill-to volume of `2`. The Volume option will be unused and the stock solution prepared according to the provided formula. WaterfallAxisFillingNotSupported The specified Filling method (`1`) is not currently supported when PlotType is set to WaterfallPlot. Defaulting Filling to Bottom. WaterfallSidePlotMissing The requested waterfall reference plot `1` could not be resolved from the input data. The waterfall plot will be generated with no reference. WavelengthMismatch Wavelength option does not match the wavelengh of ReferenceField Chromatogram, default to `1`. WavelengthRequired To plot the data as a ListLinePlot a specific wavelength must be requested. WavelengthResolutionAdjusted The following wavelength resolution options, `1`, can only be specific values and were set to the closest one. WavelengthsSwapped Some of the emission wavelengths are smaller than their corresponding excitation wavelengths. You may wish to double check these specifications. WavelengthUnavailable No data is available at the requested wavelength. Instead the data for `1` will be used. WesConcentratedLoadingBufferNotOptimal The user-supplied ConcentratedLoadingBuffer option, `1`, is not optimal for the MolecularWeightRange option, `2`. Use of a non-default ConcentratedLoadingBuffer will lead to instrument and data miscalibration. Please consider allowing the ConcentratedLoadingBuffer option to automatically resolve. WesHighDynamicRangeImagingNotPossible Since the user-supplied SignalDetectionTimes, `1`, are not the default values, High Dynamic Range (HDR) image processing is not possible. If this is not desired, please leave the SignalDetectionTimes as their default values, {1*Second,2*Second,4*Second,8*Second,16*Second,32*Second,64*Second,128*Second,512*Second}. WesInputsShouldBeDiluted The following inputs, `1`, have TotalProteinConcentrations greater than 3 mg/mL and are not being diluted with the aliquot sample preparation options. It is highly recommended that lysate samples be diluted with Model[Sample,StockSolution,"Simple Western 0.1X Sample Buffer"] to a TotalProteinConcentration below 2-3 mg/mL, or the amount of sample loaded into the capillary will be too large and data will be adversely affected. WesLadderNotOptimalForMolecularWeightRange The user-supplied Ladder option, `1`, is not optimal for the MolecularWeightRange option, `2`. Use of a non-default Ladder will lead to instrument and data miscalibration. Please consider allowing the Ladder option to automatically resolve. WesternPrimaryAntibodyVolumeLarge The following resolved or user-supplied PrimaryAntibodyVolumes, `1`, are 4 uL or larger. To conserve PrimaryAntibody, consider lowering the PrimaryAntibodyVolume and PrimaryAntibodyDiluentVolumes, `2`, while preserving the ratio between them. Alternatively, consider using the PreparatoryPrimitives option to pre-dilute the PrimaryAntibody to the desired concentration, and set the PrimaryAntibodyDilutionFactor to 1. WesternSecondaryAntibodyVolumeLow The sum of the SecondaryAntibodyVolume and StandardSecondaryAntibodyVolume, `1`, is less than the ideal 10 uL for the following pairs of input samples and antibodies, `2`. Please consider letting the SecondaryAntibodyVolume and StandardSecondaryAntibodyVolume options automatically resolve, or making sure that the sum of these two volumes is at least 10 uL. WesternStandardPrimaryAntibodyVolumeRatioNonIdeal For the following pairs of input samples and antibodies, `1`, the StandardPrimaryAntibodyVolume, `2`, is not between 7-13% of the sum of the PrimaryAntibodyVolume, PrimaryAntibodyDiluentVolume, and the StandardPrimaryAntibodyVolume, `3`. Please consider letting these options automatically resolve. WesTotalProteinConcentrationNotInformed The following primary input, `1`, does not have TotalProteinConcentration informed. TotalProteinConcentration is required to determine if the input samples are too concentrated. WesWashBufferNotOptimal The user-supplied WashBuffer option, `1`, is not optimal for the Experiment and may lead to non-optimal labeling and signal detection. Please consider using the default WashBuffer, Model[Sample,"Simple Western Wash Buffer"]. WetAndPressure Pressure application was requested for `1` sample(s), which will be liquid/wetted upon measurement. Pressure application may adversely affect measurement, but will continue. WetAndPressureBlanks Pressure application was requested for `1` Blanks, which are/is liquid. Pressure application may adversely affect measurement, but will continue. ZeroAmountManipulationRemoved The manipulation `1` has been removed from the manipulation list as it contained all 0 amounts. ZeroAmountRemoved The manipulation `1` includes an amount of 0. This mass-transfer has been removed from the manipulation.
Input
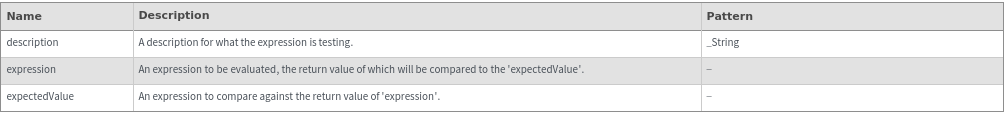
Output

General Options