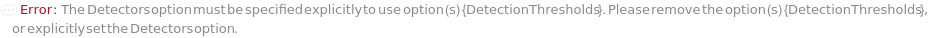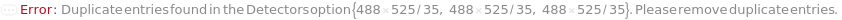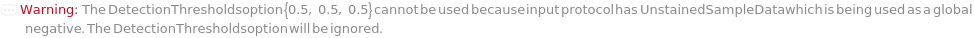AnalyzeCompensationMatrix
AnalyzeCompensationMatrix[FlowCytometryProtocol]⟹CompensationMatrix
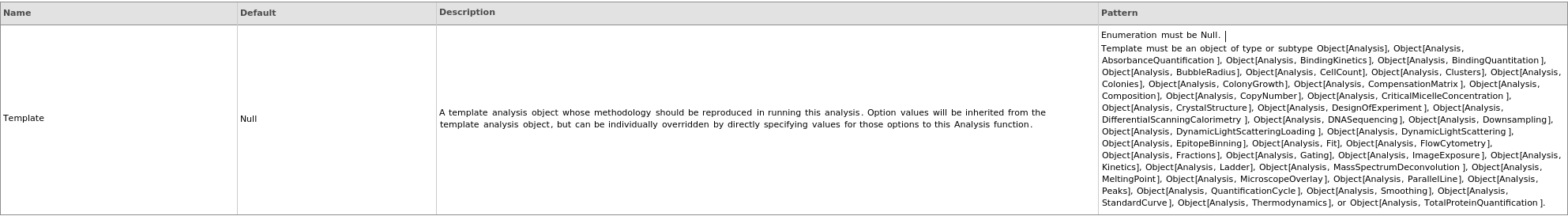
uses the AdjustmentSampleData collected in the provided FlowCytometryProtocol to compute the CompensationMatrix that corrects for fluorophore signal overlap.
Examples
open allclose allBasic Examples (5)
Compute a compensation matrix for each flow cytometry protocol in a list:
Compute a compensation matrix from a flow cytometry protocol with CompensationSamplesIncluded of True. The compensation matrix is used to correct for fluorophore spillover between detectors:
If the input protocol contains an UnstainedSample, it will be used as a universal negative. Thresholds will not be used because populations will be clearly defined. The preview will show the separation between the positive and negative populations for each compensation sample:
If the input protocol contains CompensationSamples which contain both positive and negative controls, the command builder preview will generate an interactive app which can be used to set the thresholds between positive and negative populations. Please load the function call in the builder to use this app:
Retrieve the numerical compensation matrix from a CompensationMatrix analysis object:
Options (5)
DetectionLabels (1)
DetectionThresholds (2)
If input protocol does not use an unstained global negative, for each detector (and corresponding AdjustmentSampleData), specify the signal threshold above which a cell will be considered to be positive for the label assigned to that detector:
If the input protocol uses an unstained sample as a global negative (i.e. UnstainedSampleData is informed), then DetectionThresholds will be ignored:
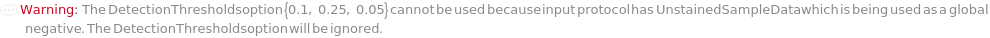
Detectors (1)
Messages (8)
AdjustmentDataNotFound (1)
Warning shows if input protocol has AdjustmentSamplesIncluded equal to True, but one or more AdjustmentSamples (index-matched to Detectors) do not have a corresponding data object in AdjustmentSampleData. Analysis will proceed, and the detectors missing AdjustmentSampleData will be assumed not to spillover into other detectors: