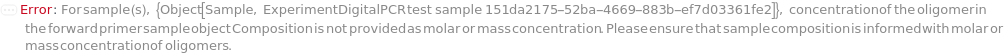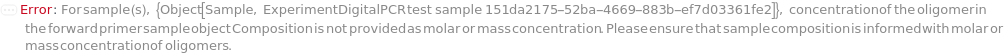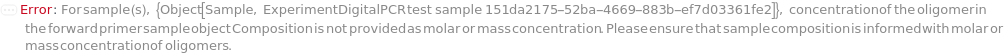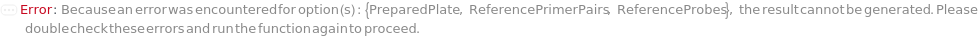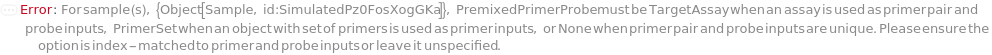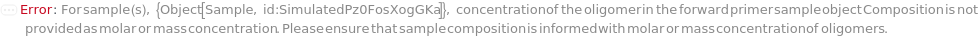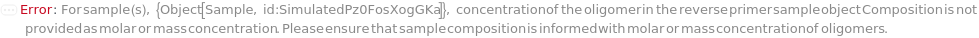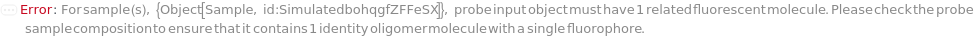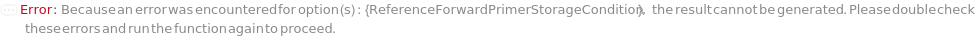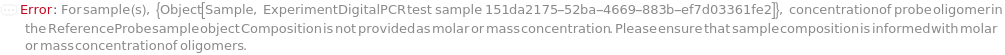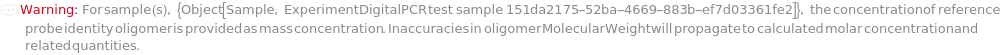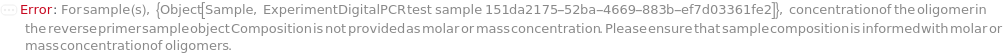General
Instrument
The device that takes a reaction mixture with target DNA or RNA, primers, probes and master mix, partitions them into nanoliter droplets, amplifies the nucleic acid templates, detects fluorescence signals from the droplets and quantifies the copy number of DNA or RNA targets.
Default Value: Model[Instrument, Thermocycler, DigitalPCR, Bio-Rad QX One]
Pattern Description: An object of type or subtype Model[Instrument, Thermocycler, DigitalPCR] or Object[Instrument, Thermocycler, DigitalPCR]
Programmatic Pattern: ObjectP[{Model[Instrument, Thermocycler, DigitalPCR], Object[Instrument, Thermocycler, DigitalPCR]}]
PreparedPlate
Indicates if the input with samples, primers, probes and master mix is already loaded on a DropletCartridge and ready to perform digital PCR without further reagent addition.
Default Calculation: Automatically resolves to True if the input samples are in a container matching "Model[Container,Plate,DropletCartridge,"Bio-Rad GCR96 Digital PCR Cartridge"]", and no PrimerPair and Probe inputs are specified, otherwise resolves to False.
Pattern Description: True or False.
Programmatic Pattern: BooleanP | Automatic
DropletCartridge
The plate with integrated microfluidics used to partition samples into droplets. Each microfluidic unit contains a sample well, channels for the sample, oil and vacuum lines to facilitate the formation of nanoliter size droplets, and a well to collect the generated droplets.
Default Value: Model[Container, Plate, DropletCartridge, Bio-Rad GCR96 Digital PCR Cartridge]
Pattern Description: An object of type or subtype Model[Container, Plate, DropletCartridge] or list of one or more an object of type or subtype Object[Container, Plate, DropletCartridge] or a prepared sample entries.
Programmatic Pattern: ObjectP[Model[Container, Plate, DropletCartridge]] | {(ObjectP[Object[Container, Plate, DropletCartridge]] | _String)..}
DropletGeneratorOil
The immiscible liquid that will be used to generate nanoliter-sized aqueous droplets in microfluidic channels of DropletCartridge.
Default Value: Model[Sample, QX One Droplet Generation Oil]
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample.
Programmatic Pattern: ObjectP[{Model[Sample], Object[Sample]}] | _String
DropletReaderOil
The immiscible liquid that will be used to separate PCR amplified droplets and facilitate fluorescence signal detection from individual droplets.
Default Value: Model[Sample, QX One Droplet Reader Oil]
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample.
Programmatic Pattern: ObjectP[{Model[Sample], Object[Sample]}] | _String
AmplitudeMultiplexing
Number of genetic targets to be detected in each fluorescence channel. Multiple genetic targets can be detected in a single channel using gradations of different primer and probe concentrations to control amplification of the gene and fluorescence signal amplitude of droplets.
Default Calculation: Automatically tallies the number of amplification targets in each fluorescence channel using the fluorophore emission wavelength of target probe(s) and reference probe(s). If any channel has more than one target and/or reference, resolves to a list of number of targets per channel; otherwise resolves to Null.
Pattern Description: List of one or more {Wavelength, Multiplexed Targets} entries or Null.
Programmatic Pattern: ({{dPCREmissionWavelengthP, GreaterEqualP[0, 1]}..} | Automatic) | Null
Index Matches to: experiment samples
NumberOfReplicates
Number of times each of the input samples should be analyzed using identical experimental parameters.
Pattern Description: Greater than or equal to 2 in increments of 1 or Null.
Programmatic Pattern: GreaterEqualP[2, 1] | Null
Sample Preparation
SampleVolume
For each sample, the portion of the reaction volume that is made up of the sample from which the target is amplified.
Default Calculation: Automatically resolves to 0
μL when PreparedPlate is True, otherwise resolves to 4
μL.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 100 microliters.
Programmatic Pattern: RangeP[0*Microliter, 100*Microliter] | Automatic
Index Matches to: experiment samples
PremixedPrimerProbe
For each sample, indicates if primerPair and probe input has an assay that contains a set of primers and a probe or if a primerPair input contains a primer set or if all three index-matched inputs are separate objects. The primerPair and probe inputs should have the same object of type Object[Sample] or Model[Sample]. Volume of the pre-mixed assay is calculated from the concentration of the probe oligomer in the mixture.
Default Calculation: Automatically resolves to TargetAssay if the primerPair and probe inputs have the same object, resolves to PrimerSet if only primerPair inputs have the same object, resolves to None if primerPair and probe have unique objects and resolves to Null if primerPairs and probes inputs are Null.
Pattern Description: List of one or more TargetAssay, PrimerSet, or None entries or Null.
Programmatic Pattern: ({(TargetAssay | PrimerSet | None)..} | Automatic) | Null
Index Matches to: experiment samples
ForwardPrimerConcentration
For each sample, the molarity of the short oligomer designed to bind the antisense strand of a template sequence. A primer functions as an anchor for polymerases and transcriptases during polymerase chain reaction.
Default Calculation: Automatically resolves to match ReversePrimerConcentration; otherwise calculated using the formula: ForwardPrimerConcentration=(max concentration)*Table[(N-t)/N,{t,0,T-1}], where (max concentration)=1350 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); or resolves to 900 nM for forward primer in channels with a single amplification product.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 1500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nanomolar, 1500*Nanomolar]..} | Automatic) | Null
Index Matches to: experiment samples
ForwardPrimerVolume
For each sample, the volume of forward primer(s) to be added to the reaction.
Default Calculation: Automatically calculated using the formula: ForwardPrimerVolume=(ForwardPrimerConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
ReversePrimerConcentration
For each sample, the molarity of the short oligomer designed to bind the sense (coding) strand of a template sequence. A primer functions as an anchor for polymerases and transcriptases during polymerase chain reaction.
Default Calculation: Automatically resolves to match ForwardPrimerConcentration; otherwise calculated using the formula: ReversePrimerConcentration=(max concentration)*Table[(N-t)/N,{t,0,T-1}], where (max concentration)=1350 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); or resolves to 900 nM for reverse primer in channels with a single amplification product.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 1500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nano*Molar, 1500*Nano*Molar]..} | Automatic) | Null
Index Matches to: experiment samples
ReversePrimerVolume
For each sample, the volume of reverse primer(s) to be added to the reaction.
Default Calculation: Automatically calculated using the formula: ReversePrimerConcentration=(ReversePrimerConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
ProbeConcentration
For each sample, the molarity of the short oligomer strand containing a reporter and quencher, and designed to be complementary to a template sequence being amplified. Fluorescence signal is observed when the probe is hydrolyzed by a polymerase and the fluorophore is released.
Default Calculation: Automatically calculated using the formula: ProbeConcentration=(max concentration)*Table[(N-t)/N,{t,0,T-1}], where (max concentration)=375 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); otherwise resolves to 250 nM for reverse primer in channels with a single amplification product.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nano*Molar, 500*Nano*Molar]..} | Automatic) | Null
Index Matches to: experiment samples
ProbeVolume
For each sample, the volume of probe(s) to be added to the reaction.
Default Calculation: Automatically resolved using the formula: ProbeVolume=(ProbeConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
ReferencePrimerPairs
For each sample, the pair of short oligomer strands designed to amplify a reference gene (such as a housekeeping gene), whose expression should not be different between samples. Primers can be specified as individual objects for forward and reverse primers. Primer-sets and pre-mixed assays with primers and probes can also be used.
Pattern Description: List of one or more {Reference Forward Primer, Reference Reverse Primer} entries or {Reference Forward Primer, Reference Reverse Primer} or Null.
Programmatic Pattern: ({{ObjectP[{Model[Sample], Object[Sample]}] | _String, ObjectP[{Model[Sample], Object[Sample]}] | _String}..} | {Null, Null}) | Null
Index Matches to: experiment samples
ReferencePremixedPrimerProbe
For each sample, indicates if ReferencePrimerPairs and ReferenceProbes have assays that contains a set of primers and a probe or if a ReferencePrimerPairs input contains a primer set or if all three index-matched reference objects are separate. The ReferencePrimerPairs and ReferenceProbes should have the same object of type Object[Sample] or Model[Sample]. Volume of the pre-mixed assay is calculated from the concentration of the probe oligomer in the mixture.
Default Calculation: Automatically resolves to TargetAssay if the reference primer pair and reference probe inputs have the same object, resolves to PrimerSet if only reference primer pair inputs have the same object, resolves to None if reference primer pair and reference probe have unique objects and resolves to Null if reference primer pairs and probes are Null.
Pattern Description: List of one or more TargetAssay, PrimerSet, or None entries or Null.
Programmatic Pattern: ({(TargetAssay | PrimerSet | None)..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceForwardPrimerConcentration
For each sample, the molarity of reference gene(s) specific oligomer that binds to the antisense strand of the template.
Default Calculation: Automatically resolves to match ReferenceReversePrimerConcentration; otherwise calculated using the formula: ReferenceForwardPrimerConcentration=(max concentration)*Table[(N-t)/N,{t,T,N-1}], where (max concentration)=1350 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); or resolves to 900 nM for forward primer in channels with a single amplification product or Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 1500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nano*Molar, 1500*Nano*Molar]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceForwardPrimerVolume
For each sample, the volume of forward primer(s) for reference gene(s) that will be added to the reaction.
Default Calculation: Automatically calculated using the formula: ReferenceForwardPrimerConcentration=(ReferenceForwardPrimerConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object; Resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceReversePrimerConcentration
For each sample, the molarity of reference gene(s) specific oligomer that binds to the sense (coding) strand of the template.
Default Calculation: Automatically resolves to match ReferenceForwardPrimerConcentration; otherwise calculated using the formula: ReferenceReversePrimerConcentration=(max concentration)*Table[(N-t)/N,{t,T,N-1}], where (max concentration)=1350 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); or resolves to 900 nM for reverse primer in channels with a single amplification product or Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 1500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nano*Molar, 1500*Nano*Molar]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceReversePrimerVolume
For each sample, the volume of reverse primer(s) for reference gene(s) that will be added to the reaction.
Default Calculation: Automatically calculated using the formula: ReferenceReversePrimerConcentration=(ReferenceReversePrimerConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object; Resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceProbes
For each sample, the short oligomer strand containing a fluorescence reporter and quencher, which enhances its fluorescence upon separation from the quencher.
Pattern Description: List of one or more an object of type or subtype Model[Sample] or Object[Sample] or a prepared sample entries or Null.
Programmatic Pattern: {(ObjectP[{Model[Sample], Object[Sample]}] | _String)..} | Null
Index Matches to: experiment samples
ReferenceProbeConcentration
For each sample, the molarity of probe(s) for reference gene(s) in the reaction.
Default Calculation: Automatically calculated using the formula: ReferenceProbeConcentration=(max concentration)*Table[(N-t)/N,{t,T,N-1}], where (max concentration)=375 Nanomolar, N=(number of targets + number of references) in a channel resolved in AmplitudeMultiplexing, and T=(number of targets); otherwise resolves to 250 nM for probes in channels with a single amplification product or Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 nanomolar and less than or equal to 500 nanomolar entries or Null.
Programmatic Pattern: ({RangeP[0*Nano*Molar, 500*Nano*Molar]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceProbeVolume
For each sample, the volume of the probe(s) for reference gene(s) that will be added to the reaction.
Default Calculation: Automatically resolved using the formula: ProbeVolume=(ProbeConcentration*ReactionVolume)/(stock concentration), where stock concentration is downloaded from the compositional information of the corresponding oligomer object; resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more greater than or equal to 0 microliters and less than or equal to 100 microliters entries or Null.
Programmatic Pattern: ({RangeP[0*Microliter, 100*Microliter]..} | Automatic) | Null
Index Matches to: experiment samples
MasterMix
For each sample, the stock solution composed of enzymes (DNA polymerase and optionally reverse transcriptase), buffer and nucleotides used to amplify DNA or RNA targets in the sample. It is recommended to select the MasterMix even if it is in the sample as the type of master mix can affect the size of droplets generated and selection of droplets for signal detection.
Default Calculation: Automatically resolves to Model[Sample,"Bio-Rad One-Step RT-ddPCR Advanced Kit for Probes, Buffer"] if input samples contain RNA templates or if any reverse transcription options are informed. The RT buffer requires addition of reverse transcriptase and DTT, so the master mix will be automatically replaced with Model[Sample,StockSolution,"Bio-Rad One-Step RT-ddPCR Advanced Kit for Probes, 2x Master Mix"] in the MasterMixes field of Object[Protocol,DigitalPCR]. Resolves to Model[Sample,"Bio-Rad ddPCR Multiplex Supermix"] if multiple probes are used, otherwise resolves to Model[Sample,"Bio-Rad ddPCR Supermix for Probes (No dUTP)"].
Pattern Description: An object of type or subtype Object[Sample] or Model[Sample] or a prepared sample.
Programmatic Pattern: (ObjectP[{Object[Sample], Model[Sample]}] | _String) | Automatic
Index Matches to: experiment samples
MasterMixConcentrationFactor
For each sample, the amount by which the MasterMix stock solution must be diluted with Diluent in order to achieve 1X buffer concentration in the reaction.
Default Calculation: Automatically resolves to Null if PreparedPlate is True, otherwise set to ConcentratedBufferDilutionFactor from Model[Sample] of MasterMix.
Pattern Description: Greater than 0. in increments of 1 or Null.
Programmatic Pattern: (GreaterP[0., 1] | Automatic) | Null
Index Matches to: experiment samples
MasterMixVolume
For each sample, the volume of the stock solution (composed of enzymes (DNA polymerase and optionally reverse transcriptase), buffer and nucleotides) that will be added to the sample.
Default Calculation: Automatically resolves to Null if PreparedPlate is True, otherwise set to MasterMixVolume=ReactionVolume/MasterMixConcentrationFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 100 microliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, 100*Microliter] | Automatic) | Null
Index Matches to: experiment samples
Diluent
For each sample, stock solution used to reach ReactionVolume, once all other components have been added.
Default Value: Model[Sample, Nuclease-free Water]
Pattern Description: An object of type or subtype Object[Sample] or Model[Sample] or a prepared sample.
Programmatic Pattern: ObjectP[{Object[Sample], Model[Sample]}] | _String
Index Matches to: experiment samples
DiluentVolume
For each sample, the volume of stock solution to be added to the reaction.
Default Calculation: Automatically set according to the equation DiluentVolume=ReactionVolume-(SampleVolume+ForwardPrimerVolume+ReversePrimerVolume+ProbeVolume+ReferenceForwardPrimerVolume+ReferenceReversePrimerVolume+ReferenceProbeVolume+MasterMixVolume) if Diluent is not set to Null, or Null otherwise.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 100 microliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, 100*Microliter] | Automatic) | Null
Index Matches to: experiment samples
ReactionVolume
For each sample, the volume of reaction including sample, primers, probes, master mix and buffer. A LoadingVolume of 21 Microliter will be transferred to DropletCartridge well after reaction components are mixed.
Default Value: 40 microliters
Pattern Description: Greater than or equal to 30 microliters and less than or equal to 100 microliters.
Programmatic Pattern: RangeP[30*Microliter, 100*Microliter]
Index Matches to: experiment samples
ActiveWell
The well in which the reaction will be conducted. The position can be designated with a well index when a single DropletCartridge is in use. An alternate input with plate index and well index can be used to configure multiple DropletCartridge plates in a single experiment. Samples can be distributed in up to 5 individual 96-well DropletCartridges.
Default Calculation: Automatically resolves the well indices. For variable thermocycling conditions, samples are grouped with compatible conditions. Samples containing RNA are prioritized for droplet generation and thermocycling.
Pattern Description: List of one or more a string that matches the pattern: WellP entries or list of one or more {Plate Index, Well Index} entries.
Programmatic Pattern: ({WellP..} | {{1 | 2 | 3 | 4 | 5, WellP}..}) | Automatic
PassiveWells
Droplet generation is conducted in parallel on a section of 16 sample units in a DropletCartridge. Each section is composed of an adjacent pair of an odd and an even numbered column such as {1,2}. Unused wells in a section are filled with viscosity matched control buffer to enable parallel processing and no data is generated from these wells.
Default Calculation: Automatically resolves the well indices for empty wells within a 16 unit section that has 1 or more samples. Resolves to None if each 16 unit section in use is filled with samples. Sections without any samples are skipped during the assay.
Pattern Description: List of one or more a string that matches the pattern: WellP entries or list of one or more {Plate Index, Well Index} entries or None.
Programmatic Pattern: ({WellP..} | {{1 | 2 | 3 | 4 | 5, WellP}..} | None) | Automatic
PassiveWellBuffer
Control solution with viscosity matched to master mix. 20 Microliter of buffer is added to PassiveWells.
Default Value: Model[Sample, Bio-Rad ddPCR Buffer Control for Probes]
Pattern Description: An object of type or subtype Object[Sample] or Model[Sample] or a prepared sample.
Programmatic Pattern: ObjectP[{Object[Sample], Model[Sample]}] | _String
Sample Dilution
SampleDilution
For each sample, indicates if Diluent should be used to generate a series of decreasing target concentrations to ensure that an unknown sample falls within the DigitalPCR detection range of 0.25 to 5,000 targets/
μL.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
SerialDilutionCurve
For each sample, the collection of dilutions that are performed before the SampleVolume is transferred to a mixture of primers, probes, MasterMix and Diluent. For Serial Dilution Factors, the sample will be diluted with the Diluent by the dilution factor at each transfer step. For example, a SerialDilutionCurve of {36*Microliter,{1,0.8,0.5,0.5}} for a 100 mg/mL sample will result in 4 dilutions with concentrations of 100 mg/mL, 80 mg/mL, 40 mg/mL, and 20 mg/mL. For Serial Dilution Volumes, the Transfer Volume is taken out of the sample and added to a second well with the Diluent Volume of the Diluent. It is mixed, then the Transfer Volume is taken out of that well to be added to a third well. This is repeated to make Number Of Dilutions diluted samples. For example, if a 100 mg/mL sample with a Transfer Volume of 30 Microliters, a Diluent Volume of 10 Microliters and a Number of Dilutions of 2 is used, it will create a DilutionCurve of 75 mg/mL, 56.3 mg/mL, and 42.2 mg/mL.
Default Calculation: Automatically resolves to {50*Microliter,{0.1,4}}, where 50
μL is the sample volume, 0.1 is the constant dilution factor and 4 is the number of dilutions, when SampleDilution is True, otherwise resolves to Null.
Pattern Description: Serial Dilution Factor or Serial Dilution Volumes or Null.
Programmatic Pattern: (({GreaterP[0*Microliter], GreaterEqualP[0*Microliter], GreaterP[1, 1]} | {GreaterEqualP[10*Microliter], {RangeP[0, 1], GreaterP[1, 1]} | {RangeP[0, 1]..}}) | Automatic) | Null
Index Matches to: experiment samples
DilutionMixVolume
For each sample, the volume that is pipetted in and out to mix the sample and the Diluent.
Default Calculation: Automatically resolves to 10
μL if SampleDilution is True, otherwise resolves to Null.
Pattern Description: Greater than or equal to 0 microliters or Null.
Programmatic Pattern: (GreaterEqualP[0*Microliter] | Automatic) | Null
Index Matches to: experiment samples
DilutionNumberOfMixes
For each sample, the number of cycles of pipetting in and out that is used to mix the sample and the Diluent.
Default Calculation: Automatically resolves to 10 if SampleDilution is True, otherwise resolves to Null.
Pattern Description: Greater than or equal to 0 and less than or equal to 20 in increments of 1 or Null.
Programmatic Pattern: (RangeP[0, 20, 1] | Automatic) | Null
Index Matches to: experiment samples
DilutionMixRate
For each sample, the speed at which the DilutionMixVolume is pipetted in and out to mix the sample and the Diluent.
Default Calculation: Automatically resolves to 30
μL/s if SampleDilution is True, otherwise resolves to Null.
Pattern Description: Greater than or equal to 0.4 microliters per second and less than or equal to 250 microliters per second or Null.
Programmatic Pattern: (RangeP[0.4*(Microliter/Second), 250*(Microliter/Second)] | Automatic) | Null
Index Matches to: experiment samples
Plate Seal
PlateSealInstrument
The instrument used to apply heat-seal digital PCR plates with foil prior to assay run.
Default Value: Model[Instrument, PlateSealer, Bio-Rad PX1 Plate Sealer]
Pattern Description: An object of type or subtype Model[Instrument, PlateSealer] or Object[Instrument, PlateSealer]
Programmatic Pattern: ObjectP[{Model[Instrument, PlateSealer], Object[Instrument, PlateSealer]}]
PlateSealFoil
The pierceable membrane used to seal digital PCR plate.
Default Value: Model[Item, PlateSeal, GCR96 PCR Plate Seal, Pierceable Heat-Sealed Aluminum]
Pattern Description: An object of type or subtype Model[Item] or Object[Item]
Programmatic Pattern: ObjectP[{Model[Item], Object[Item]}]
PlateSealTemperature
The temperature that will be used to heat the foil for sealing a plate.
Default Value: 180 degrees Celsius
Pattern Description: Greater than or equal to 100 degrees Celsius and less than or equal to 190 degrees Celsius.
Programmatic Pattern: RangeP[100*Celsius, 190*Celsius]
PlateSealTime
The duration of time used for applying PlateSealTemperature to seal the digital PCR plate.
Default Value: 0.5 seconds
Pattern Description: Greater than or equal to 0.5 seconds and less than or equal to 10 seconds.
Programmatic Pattern: RangeP[0.5*Second, 10*Second]
Reverse Transcription
ReverseTranscription
For each sample, indicates if one-step reverse transcription is performed in order to convert RNA input samples to cDNA. Thermocycling is performed after samples are partitioned into droplets and collected in the droplet well of their corresponding microfluidic unit. All sample emulsions on a DropletCartridge are thermocycled simultaneously.
Default Calculation: Automatically set to True if any reverse transcription related options are set, and False if none are set.
Pattern Description: True or False.
Programmatic Pattern: BooleanP | Automatic
Index Matches to: experiment samples
ReverseTranscriptionTime
For each sample, the length of time for which the reaction volume will held at ReverseTranscriptionTemperature in order to convert RNA to cDNA.
Default Calculation: Automatically set to 15 minutes if ReverseTranscription is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 2 hours or Null.
Programmatic Pattern: (RangeP[0*Second, 2*Hour] | Automatic) | Null
Index Matches to: experiment samples
ReverseTranscriptionTemperature
For each sample, the initial temperature at which the sample should be heated to in order to activate the reverse transcription enzymes in the master mix to convert RNA to cDNA.
Default Calculation: Automatically set to 50
°Celsius if ReverseTranscription is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[4*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
ReverseTranscriptionRampRate
For each sample, the rate at which the reaction volume is heated to bring the sample to the reverse transcription temperature.
Default Calculation: Automatically set to 1.6
°Celsius/Second if ReverseTranscription is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second or Null.
Programmatic Pattern: (RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)] | Automatic) | Null
Index Matches to: experiment samples
Polymerase Activation
Activation
For each sample, indicates if polymerase hot start activation is performed. In order to reduce non-specific amplification, modern enzymes can be made room temperature stable by inhibiting their activity via thermolabile conjugates. Once an experiment is ready to be run, this inhibition is removed by heating the reaction to ActivationTemperature. The activation step is recommended as ActivationTemperature promotes stabilization of droplet boundary through a reaction between MasterMix and DropletGeneratorOil prior to thermocycling.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
ActivationTime
For each sample, the length of time for which the reaction volume is held at ActivationTemperature in order to remove the thermolabile conjugates inhibiting enzyme activity.
Default Calculation: Automatically set to 10 minutes if Activation is set to True.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 2 hours or Null.
Programmatic Pattern: (RangeP[0*Second, 2*Hour] | Automatic) | Null
Index Matches to: experiment samples
ActivationTemperature
For each sample, the temperature at which at which the thermolabile conjugates inhibiting enzyme activity are removed.
Default Calculation: Automatically set to 95
°Celsius if Activation is set to True.
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[4*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
ActivationRampRate
For each sample, the rate at which the reaction is heated to bring the sample to the ActivationTemperature.
Default Calculation: Automatically set to 1.6
°Celsius/Second if Activation is set to True.
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second or Null.
Programmatic Pattern: (RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)] | Automatic) | Null
Index Matches to: experiment samples
Denaturation
DenaturationTime
For each sample, the length of time for which the reaction is held at DenaturationTemperature in order to separate any double stranded template DNA into single strands.
Default Value: 30 seconds
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 2 hours.
Programmatic Pattern: RangeP[0*Second, 2*Hour]
Index Matches to: experiment samples
DenaturationTemperature
For each sample, the temperature to which the sample is heated in order to disassociate double stranded template DNA into single strands.
Default Value: 95 degrees Celsius
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 100 degrees Celsius.
Programmatic Pattern: RangeP[4*Celsius, 100*Celsius]
Index Matches to: experiment samples
DenaturationRampRate
For each sample, the rate at which the reaction is heated to bring the sample to DenaturationTemperature.
Default Value: 1.8 degrees Celsius per second
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second.
Programmatic Pattern: RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)]
Index Matches to: experiment samples
Primer Annealing
PrimerAnnealing
For each sample, indicates if annealing should be performed. Lowering the temperature during annealing allows primers to bind to template DNA to serve as anchor points for DNA polymerization. It is highly recommended to use PrimerAnnealing as a single step to perform annealing and extension if the working temperature of the polymerase in MasterMix is 60
°Celsius.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
PrimerAnnealingTime
For each sample, the length of time for which the reaction is held at PrimerAnnealingTemperature in order to allow binding of primers to the template DNA.
Default Calculation: Automatically set to 60 seconds if PrimerAnnealing or PrimerGradientAnnealing is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 2 hours or Null.
Programmatic Pattern: (RangeP[0*Second, 2*Hour] | Automatic) | Null
Index Matches to: experiment samples
PrimerAnnealingTemperature
For each sample, the temperature to which the sample is heated in order to allow primers to bind to the template strands.
Default Calculation: Automatically set to 60
°Celsius if PrimerAnnealing is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[4*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
PrimerAnnealingRampRate
For each sample, the rate at which the reaction is heated to bring the sample to PrimerAnnealingTemperature.
Default Calculation: Automatically set to 2.0
°Celsius/Second if PrimerAnnealing is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second or Null.
Programmatic Pattern: (RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)] | Automatic) | Null
Index Matches to: experiment samples
Primer Gradient Annealing
PrimerGradientAnnealing
For each sample, indicates if a gradation of temperature should be used for annealing across columns such that each row has the same annealing temperature. A linearly decreasing series of temperatures for 8 rows is calculated as follows: Range[Tmin,Tmax,(Tmax-Tmin)/7], where Tmin is the minimum temperature set to Row 'A' and Tmax is the maximum temperature set to Row 'H'. A temperature gradient can be used to optimize annealing by setting Tmin and Tmax to be (expected annealing temperature) +/- 5 Celsius. An aliquot of reaction mixture with the same target, primer pair and probe is added to at least one well per row of a DropletCartridge and the amplification efficiency is determined by the fluorescence signal amplitude of the droplets after PCR.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
PrimerGradientAnnealingMinTemperature
For each sample, the lower value of the temperature range that will be used to calculate annealing temperature of each row.
Default Calculation: Automatically set to 55
°Celsius if PrimerGradientAnnealing is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 30 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[30*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
PrimerGradientAnnealingMaxTemperature
For each sample, the upper value of the temperature range that will be used to calculate annealing temperature of each row.
Default Calculation: Automatically set to 65
°Celsius if PrimerGradientAnnealing is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 30 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[30*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
PrimerGradientAnnealingRow
For each sample, the tier in which the sample will be located in the DropletCartridge. "Row A" is the minimum temperature and "Row H" is the maximum temperature, while the rows in between have a gradation of temperature.
Default Calculation: Automatically resolves the row & temperature assignment for sample with PrimerGradientAnnealing set to True, where the first sample is assigned to "Row A", second sample to "Row B" and so on; otherwise resolves to Null if PrimerGradientAnnealing is False.
Pattern Description: Row A, Row B, Row C, Row D, Row E, Row F, Row G, or Row H or {Row, Temperature} or Null.
Programmatic Pattern: ((("Row A" | "Row B" | "Row C" | "Row D" | "Row E" | "Row F" | "Row G" | "Row H") | {"Row A" | "Row B" | "Row C" | "Row D" | "Row E" | "Row F" | "Row G" | "Row H", RangeP[30*Celsius, 100*Celsius]}) | Automatic) | Null
Index Matches to: experiment samples
Strand Extension
Extension
For each sample, indicates if an extension step should be performed as a separate step.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
ExtensionTime
For each sample, the length of time for which the polymerase synthesizes a new DNA strand using the provided primer pairs and template DNA.
Default Calculation: Automatically set to 60 Second if Extension is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 2 hours or Null.
Programmatic Pattern: (RangeP[0*Second, 2*Hour] | Automatic) | Null
Index Matches to: experiment samples
ExtensionTemperature
For each sample, the temperature at which the sample is held to allow the polymerase to synthesis new DNA strand from the template DNA.
Default Calculation: Automatically set to 60
°Celsius if Extension is set to True, and Null if it is set to False.
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 100 degrees Celsius or Null.
Programmatic Pattern: (RangeP[4*Celsius, 100*Celsius] | Automatic) | Null
Index Matches to: experiment samples
ExtensionRampRate
For each sample, the rate at which the reaction is heated to bring the sample to ExtensionTemperature.
Default Calculation: Automatically set to 2.0
°Celsius/Second if Extension is set to True, and Null if it is set to False
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second or Null.
Programmatic Pattern: (RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)] | Automatic) | Null
Index Matches to: experiment samples
NumberOfCycles
For each sample, the remaining number of times the samples will undergo repeated rounds of denaturation, annealing and extension to amplify targets.
Pattern Description: Greater than or equal to 1 and less than or equal to 60.
Programmatic Pattern: RangeP[1, 60]
Index Matches to: experiment samples
Polymerase Degradation
PolymeraseDegradation
For each sample, indicates if the polymerase should be degraded at PolymeraseDegradationTemperature. After thermocycling is complete, the DropletCartridge is will remain at 25 Celsius (i.e. ambient temperature). Polymerase degradation is recommended to prevent any alterations to the droplet contents prior to signal detection.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
Index Matches to: experiment samples
PolymeraseDegradationTime
For each sample, the length of time for which the sample is held at PolymeraseDegradationTemperature.
Default Calculation: Automatically set to 10 minutes if PolymeraseDegradation is set to True, or Null otherwise
Pattern Description: Greater than or equal to 1 second and less than or equal to 1 hour or Null.
Programmatic Pattern: (RangeP[1*Second, 1*Hour] | Automatic) | Null
Index Matches to: experiment samples
PolymeraseDegradationTemperature
For each sample, the temperature to which the sample is heated to degrade the polymerase.
Default Calculation: Automatically set to 98 Celsius if PolymeraseDegradation is set to True, or Null otherwise.
Pattern Description: Greater than or equal to 4 degrees Celsius and less than or equal to 105 degrees Celsius or Null.
Programmatic Pattern: (RangeP[4*Celsius, 105*Celsius] | Automatic) | Null
Index Matches to: experiment samples
PolymeraseDegradationRampRate
For each sample, the rate at which the sample is heated to reach PolymeraseDegradationTemperature.
Default Calculation: Automatically set to 2.5 degrees Celsius per second if PolymeraseDegradation is set to True, or Null otherwise.
Pattern Description: Greater than or equal to 1. degrees Celsius per second and less than or equal to 2.5 degrees Celsius per second or Null.
Programmatic Pattern: (RangeP[1.*(Celsius/Second), 2.5*(Celsius/Second)] | Automatic) | Null
Index Matches to: experiment samples
Detection
ProbeFluorophore
For each sample, the fluorescent molecule conjugated to the probe that allows for detection of amplification.
Default Calculation: Automatically set to the model of fluorophore attached to probe oligomer.
Pattern Description: List of one or more an object of type or subtype Model[Molecule] entries.
Programmatic Pattern: {ObjectP[Model[Molecule]]..} | Automatic
Index Matches to: experiment samples
ProbeExcitationWavelength
For each sample, the wavelength of light used to excite the reporter component of the probe.
Default Calculation: Automatically set to FluorescenceExcitationMaximums of the fluorophore attached to the probe.
Pattern Description: List of one or more 495. nanometers, 535. nanometers, 538. nanometers, 647. nanometers, or 675. nanometers entries.
Programmatic Pattern: {dPCRExcitationWavelengthP..} | Automatic
Index Matches to: experiment samples
ProbeEmissionWavelength
For each sample, the wavelength of light emitted by the reporter once it has been excited.
Default Calculation: Automatically set to FluorescenceEmissionMaximums of the fluorophore attached to the probe.
Pattern Description: List of one or more 517. nanometers, 554. nanometers, 556. nanometers, 665. nanometers, or 694. nanometers entries.
Programmatic Pattern: {dPCREmissionWavelengthP..} | Automatic
Index Matches to: experiment samples
ReferenceProbeFluorophore
The fluorescent molecule conjugated to the reference probe that allows for detection of amplification.
Default Calculation: Automatically set to the model of fluorophore attached to the reference probe oligomer; resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more an object of type or subtype Model[Molecule] entries or Null.
Programmatic Pattern: ({ObjectP[Model[Molecule]]..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceProbeExcitationWavelength
The wavelength of light used to excite the reporter component of the reference probe.
Default Calculation: Automatically set to FluorescenceExcitationMaximums of the fluorophore attached to the reference probe; resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more 495. nanometers, 535. nanometers, 538. nanometers, 647. nanometers, or 675. nanometers entries or Null.
Programmatic Pattern: ({dPCRExcitationWavelengthP..} | Automatic) | Null
Index Matches to: experiment samples
ReferenceProbeEmissionWavelength
The wavelength of light emitted by the reporter once it has been excited and read by the instrument.
Default Calculation: Automatically set to FluorescenceEmissionMaximums of the fluorophore attached to the reference probe; resolves to Null if ReferencePrimerPairs is Null.
Pattern Description: List of one or more 517. nanometers, 554. nanometers, 556. nanometers, 665. nanometers, or 694. nanometers entries or Null.
Programmatic Pattern: ({dPCREmissionWavelengthP..} | Automatic) | Null
Index Matches to: experiment samples
Post Experiment
ForwardPrimerStorageCondition
For each sample, the non-default conditions under which the forward primers of this experiment should be stored after the protocol is completed. If left unset, the forward primers will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ReversePrimerStorageCondition
For each sample, the non-default conditions under which the reverse primers of this experiment should be stored after the protocol is completed. If left unset, the reverse primers will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ProbeStorageCondition
For each sample, the non-default conditions under which the probes of this experiment should be stored after the protocol is completed. If left unset, the probes will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ReferenceForwardPrimerStorageCondition
For each sample, the non-default conditions under which the forward primers for the reference of this experiment should be stored after the protocol is completed. If left unset, the forward primers for the reference will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ReferenceReversePrimerStorageCondition
For each sample, the non-default conditions under which the reverse primers for the reference of this experiment should be stored after the protocol is completed. If left unset, the reverse primers for the reference will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ReferenceProbeStorageCondition
For each sample, the non-default conditions under which the probes for the reference of this experiment should be stored after the protocol is completed. If left unset, the probes for the reference will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
MasterMixStorageCondition
For each sample, the non-default condition under which MasterMix of this experiment should be stored after the protocol is completed. If left unset, MasterMix will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
SamplesInStorageCondition
The non-default conditions under which the SamplesIn of this experiment should be stored after the protocol is completed. If left unset, SamplesIn will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (Alternatives[SampleStorageTypeP | Disposal]) | Null
Index Matches to: experiment samples
Model Input
PreparedModelContainer
Indicates the container in which a Model[Sample] specified as input to the experiment function will be prepared.
Default Calculation: If PreparedModelAmount is set to All and when the input model has a product associated with both Amount and DefaultContainerModel populated, automatically set to the DefaultContainerModel value in the product. Otherwise set to Model[Container, Vessel, "2mL Tube"].
Pattern Description: An object of type or subtype Model[Container] or Null.
Programmatic Pattern: (ObjectP[Model[Container]] | Automatic) | Null
Index Matches to: experiment samples
PreparedModelAmount
Indicates the amount of a Model[Sample] specified as input to the experiment function that will be prepared in the PreparedModelContainer. When set to All and the input model sample is not preparable, the entire amount of the input model sample that comes from one of the Products is prepared. The selected product must have both Amount and DefaultContainerModel populated, and it must not be a KitProduct. When set to All and the input model is preparable such as water, 1 Milliliter of the input model sample is prepared.
Default Calculation: Automatically set to 1 Milliliter.
Pattern Description: All or Count or Count or Mass or Volume or Null.
Programmatic Pattern: ((RangeP[1*Microliter, 20*Liter] | RangeP[1*Milligram, 20*Kilogram] | GreaterP[0*Unit, 1*Unit] | GreaterP[0., 1.] | All) | Automatic) | Null
Index Matches to: experiment samples