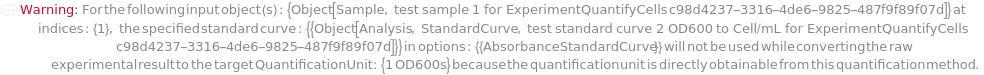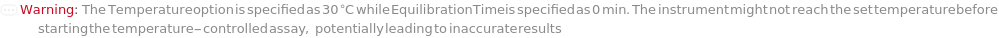General
Methods
The experimental instrumentation used to measure the cell concentration in the input cell samples. Instrumentation options include: 1) Absorbance, where the absorbance (at the provided Wavelength) of the cell sample is measured using a spectrophotometer or plate reader and then converted to QuantificationUnit using either the AbsorbanceStandardCurve or AbsorbanceStandardCoefficient; 2) Nephelometry, where the scattered light attenuation of the cell sample at 653 nm is measured using a nephelometer and then converted to QuantificationUnit using either the NephelometryStandardCurve or NephelometryStandardCoefficient. The order of methods dictates the order of operations when the protocol is executed in the lab.
Default Calculation: Automatically set to Absorbance if any Absorbance options are provided or Instruments is set to a Spectrophotometer or PlateReader. Automatically set to Nephelometry if any Nephelometry options are provided or Instruments is set to a Nephelometer. Otherwise, if QuantificationUnit is set, Methods is automatically set to the instrumentation capable of generating raw experimental results in that unit. If the QuantificationUnit cannot be directly generated by any instrumentation, Methods is automatically set to the instrumentation whose native output can be converted to the QuantificationUnit using the specified StandardCurve/StandardCoefficient, or using the standard curve found from cell models in the Composition of the input samples. Otherwise, Methods is automatically set to Absorbance.
Pattern Description: Absorbance or Nephelometry.
Programmatic Pattern: CellQuantificationMethodP | Automatic
Index Matches to: Methods
Instruments
The instrument used to measure the absorbance (at the provided Wavelength) or nephelometry (scattered light attenuation) for the input cell samples.
Default Calculation: Automatically set to include a spectrophotometer or plate reader model if Methods include Absorbance. Automatically set to include a nephelometer model if Methods include Nephelometry. Please see the documentation of ExperimentAbsorbanceIntensity and ExperimentNephelometry for more details on the how the instrument option is determined given the input cell samples and corresponding Absorbance/Nephelometry options.
Pattern Description: An object of type or subtype Model[Instrument, PlateReader], Object[Instrument, PlateReader], Model[Instrument, Spectrophotometer], Object[Instrument, Spectrophotometer], Model[Instrument, Nephelometer], or Object[Instrument, Nephelometer]
Programmatic Pattern: ObjectP[{Model[Instrument, PlateReader], Object[Instrument, PlateReader], Model[Instrument, Spectrophotometer], Object[Instrument, Spectrophotometer], Model[Instrument, Nephelometer], Object[Instrument, Nephelometer]}] | Automatic
Index Matches to: Methods
Preparation
Indicates if this unit operation is carried out primarily robotically or manually. Manual unit operations are executed by a laboratory operator and robotic unit operations are executed by a liquid handling work cell.
Pattern Description: Manual or Robotic.
Programmatic Pattern: PreparationMethodP | Automatic
WorkCell
The automated workstation with a collection of integrated instruments on which this unit operation will be will be performed if Preparation -> Robotic.
Default Calculation: Automatically set to STAR if Preparation->Robotic.
Pattern Description: STAR, bioSTAR, or microbioSTAR or Null.
Programmatic Pattern: ((STAR | bioSTAR | microbioSTAR) | Automatic) | Null
NumberOfReplicates
The number of times each of the input samples is analyzed using identical experimental parameters. If Absorbance/NephelometryAliquot is set to True, this specifies the number of replicate aliquots to prepare from each input cell sample under the same aliquot conditions.
Pattern Description: Greater than or equal to 2 in increments of 1 or Null.
Programmatic Pattern: (Null | GreaterEqualP[2, 1]) | Null
RecoupSample
Indicates if the aliquots from the corresponding input cell samples used for quantification measurement are returned to those input cell samples after the protocol is completed.
Pattern Description: True or False or Null.
Programmatic Pattern: (Null | BooleanP) | Null
Index Matches to: experiment samples
MultiMethodAliquots
Indicates if a single aliquot is taken from the input cell samples and reused in all quantification measurements (Shared) or if each measurement takes a new aliquot from the input cell samples (Individual).
Pattern Description: Shared or Individual.
Programmatic Pattern: MultiMethodAliquotsP
Post Experiment
SamplesInStorageCondition
The non-default conditions under which the SamplesIn of this experiment should be stored after the protocol is completed. If left unset, SamplesIn will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (Null | (Alternatives[SampleStorageTypeP | Disposal])) | Null
Index Matches to: experiment samples
Analysis
QuantificationUnit
The preferred unit of cell concentration to update the composition of the input sample after the quantification measurement. If the specified unit cannot be directly measured with the specified quantification Methods, Absorbance/NephelometryStandardCurve or Absorbance/NephelometryStandardCoefficient will be used to convert the raw experimental result. If the Absorbance/NephelometryStandardCurve or Absorbance/NephelometryStandardCoefficient information is neither provided nor available from the input samples' cell compositions, the unit that is directly measured with the specified quantification Methods will be used to update the composition. The final concentration used to update the composition is the mean value of any raw experimental results and any converted values from the raw experimental result in the unit of QuantificationUnit. If the QuantificationUnit is neither convertible nor directly measured by the quantification Methods, the composition of the input sample will not be updated. If the input sample contains more than one cell type, its composition will not be updated given that the quantification Methods only measure one overall quantity of all contents in the sample.
Default Calculation: Automatically set to "EmeraldCell/Milliliter" if a standard curve or standard coefficient is provided in StandardCurve/StandardCoefficient options or found from the input cell samples' cell Composition to convert the raw experimental result from Methods to "EmeraldCell/Milliliter". Otherwise, automatically set to "OD600".
Pattern Description: EmeraldCell/Milliliter or OD600.
Programmatic Pattern: CellQuantificationUnitStringP | Automatic
Index Matches to: experiment samples
AbsorbanceStandardCurve
The standard curve used to convert the raw experimental result, expressed in units of OD600 or AbsorbanceUnit, to the specified QuantificationUnit.
Default Calculation: Automatically set to the standard curve that is found from the input cell sample's cell Composition that can convert raw OD600 or AbsorbanceUnit to the target QuantificationUnit. Otherwise, automatically set to Null.
Pattern Description: An object of type or subtype Object[Analysis, StandardCurve] or Null.
Programmatic Pattern: (ObjectP[Object[Analysis, StandardCurve]] | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceStandardCoefficient
The factor used to multiply the raw experimental result in units of OD600 or AbsorbanceUnit to the specified QuantificationUnit, assuming a linear relationship.
Pattern Description: Greater than 0 or Null.
Programmatic Pattern: GreaterP[0] | Null
Index Matches to: experiment samples
NephelometryStandardCurve
The standard curve used to convert the raw experimental result, expressed in the unit of RelativeNephelometricUnit, to the specified QuantificationUnit.
Default Calculation: Automatically set to the standard curve that is found from the input cell sample's cell Composition that can convert raw RelativeNephelometricUnit to the target QuantificationUnit. Otherwise, automatically set to Null.
Pattern Description: An object of type or subtype Object[Analysis, StandardCurve] or Null.
Programmatic Pattern: (ObjectP[Object[Analysis, StandardCurve]] | Automatic) | Null
Index Matches to: experiment samples
NephelometryStandardCoefficient
The factor used to multiply the raw experimental result in units of RelativeNephelometricUnit to the specified QuantificationUnit, assuming a linear relationship.
Pattern Description: Greater than 0 or Null.
Programmatic Pattern: GreaterP[0] | Null
Index Matches to: experiment samples
Aliquoting
AbsorbanceAliquot
Indicates if an aliquot is taken from the input cell sample or the aliquot from the previous quantification measurements (if any), and is used for the subsequent Absorbance measurement. If NumberOfReplicates is specified this indicates that the input samples will also be aliquoted that number of times.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceAssayBuffer
The solution that is added to the aliquot taken from the input cell sample or the aliquot from the previous quantification measurements (if any). The volume of this solution added is the difference between the AbsorbanceAliquotAmount and the AbsorbanceAssayVolume.
Default Calculation: Automatically resolves to Model[Sample, "Milli-Q water"] if ConcentratedBuffer is not specified; otherwise, resolves to Null.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((Null | (ObjectP[{Model[Sample], Object[Sample]}] | _String)) | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceAliquotAmount
The amount of the sample that is transferred from the input cell sample or the aliquot from the previous quantification measurements (if any) into the new aliquot.
Default Calculation: Automatically set as the smaller between the current sample volume and the maximum volume of the destination container if a liquid, or the current Mass or Count if a solid or counted item, respectively.
Pattern Description: All or Volume or Null.
Programmatic Pattern: ((RangeP[1*Microliter, $CuvetteMaxVolume] | All) | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceAssayVolume
The desired total volume of the aliquoted sample plus AbsorbanceAssayBuffer.
Default Calculation: Automatically determined based on Volume and TargetConcentration option values.
Pattern Description: Greater than or equal to 1 microliter and less than or equal to 4. milliliters or Null.
Programmatic Pattern: (RangeP[1*Microliter, $CuvetteMaxVolume] | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceAliquotContainer
The desired type of container that is used to house the aliquot sample prior to the Absorbance measurement, with indices indicating grouping of samples in the same plates, if desired.
Default Calculation: Automatically set as the PreferredContainer for the AssayVolume of the sample. For plates, attempts to fill all wells of a single plate with the same model before aliquoting into the next.
Pattern Description: Container or Container with Index or list of one or more Container or Container with Index entries or Null.
Programmatic Pattern: (((ObjectP[{Model[Container], Object[Container]}] | _String) | {GreaterEqualP[1, 1], ObjectP[{Model[Container], Object[Container]}] | _String} | {((ObjectP[{Model[Container], Object[Container]}] | _String) | {GreaterEqualP[1, 1], ObjectP[{Model[Container], Object[Container]}] | _String})..}) | Automatic) | Null
AbsorbanceDestinationWell
The desired position in the corresponding AbsorbanceAliquotContainer in which the aliquot sample will be placed.
Default Calculation: Automatically resolves to A1 in containers with only one position. For plates, fills wells in the order provided by the function AllWells.
Pattern Description: Any well from A1 to H12 or list of one or more any well from A1 to H12 or any well from A1 to H12 entries or Null.
Programmatic Pattern: ((Null | (WellPositionP | {((Automatic | Null) | WellPositionP)..})) | Automatic) | Null
AbsorbanceAliquotSampleStorageCondition
The non-default conditions under which any unused aliquot sample generated for Absorbance measurement is stored after the protocol is completed.
Pattern Description: {AmbientStorage, Refrigerator, Freezer, DeepFreezer} or Disposal or Null.
Programmatic Pattern: ((PostAnalysisCellSampleStorageTypeP | Disposal) | Automatic) | Null
Index Matches to: experiment samples
NephelometryAliquot
Indicates if an aliquot is taken from the input cell sample or the aliquot from the previous quantification measurements (if any), and is used for the subsequent Nephelometry measurement. If NumberOfReplicates is specified this indicates that the input samples will also be aliquoted that number of times.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
Index Matches to: experiment samples
NephelometryAssayBuffer
The solution that is added to the aliquot taken from the input cell sample or the aliquot from the previous quantification measurements (if any). The volume of this solution added is the difference between the NephelometryAliquotAmount and the NephelometryAssayVolume.
Default Calculation: Automatically resolves to Model[Sample, "Milli-Q water"] if ConcentratedBuffer is not specified; otherwise, resolves to Null.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((Null | (ObjectP[{Model[Sample], Object[Sample]}] | _String)) | Automatic) | Null
Index Matches to: experiment samples
NephelometryAliquotAmount
The amount of the sample that is transferred from the input cell sample or the aliquot from the previous quantification measurements (if any) into the new aliquot.
Default Calculation: Automatically set as the smaller between the current sample volume and the maximum volume of the destination container if a liquid, or the current Mass or Count if a solid or counted item, respectively.
Pattern Description: All or Volume or Null.
Programmatic Pattern: ((RangeP[1*Microliter, $CuvetteMaxVolume] | All) | Automatic) | Null
Index Matches to: experiment samples
NephelometryAssayVolume
The desired total volume of the aliquoted sample plus NephelometryAssayBuffer.
Default Calculation: Automatically determined based on Volume and TargetConcentration option values.
Pattern Description: Greater than or equal to 1 microliter and less than or equal to 4. milliliters or Null.
Programmatic Pattern: (RangeP[1*Microliter, $CuvetteMaxVolume] | Automatic) | Null
Index Matches to: experiment samples
NephelometryAliquotContainer
The desired type of container that is used to house the aliquot sample prior to the Nephelometry measurement, with indices indicating grouping of samples in the same plates, if desired.
Default Calculation: Automatically set as the PreferredContainer for the AssayVolume of the sample. For plates, attempts to fill all wells of a single plate with the same model before aliquoting into the next.
Pattern Description: Container or Container with Index or list of one or more Container or Container with Index entries or Null.
Programmatic Pattern: (((ObjectP[{Model[Container], Object[Container]}] | _String) | {GreaterEqualP[1, 1], ObjectP[{Model[Container], Object[Container]}] | _String} | {((ObjectP[{Model[Container], Object[Container]}] | _String) | {GreaterEqualP[1, 1], ObjectP[{Model[Container], Object[Container]}] | _String})..}) | Automatic) | Null
NephelometryDestinationWell
The desired position in the corresponding NephelometryAliquotContainer in which the aliquot sample will be placed.
Default Calculation: Automatically resolves to A1 in containers with only one position. For plates, fills wells in the order provided by the function AllWells.
Pattern Description: Any well from A1 to H12 or list of one or more any well from A1 to H12 or any well from A1 to H12 entries or Null.
Programmatic Pattern: ((Null | (WellPositionP | {((Automatic | Null) | WellPositionP)..})) | Automatic) | Null
NephelometryAliquotSampleStorageCondition
The non-default conditions under which any unused aliquot sample generated for Nephelometry measurement is stored after the protocol is completed.
Pattern Description: {AmbientStorage, Refrigerator, Freezer, DeepFreezer} or Disposal or Null.
Programmatic Pattern: ((PostAnalysisCellSampleStorageTypeP | Disposal) | Automatic) | Null
Index Matches to: experiment samples
Blanks
AbsorbanceBlankMeasurement
Indicates if blank samples are prepared and measured prior to Absorbance measurement. The absorbance (at the provided Wavelength) of the blank samples are measured and background subtracted to account for any background signals.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
AbsorbanceBlank
The source used to generate a blank sample used to account for the background signal prior to the Absorbance measurement of the cell samples of interest.
Default Calculation: Automatically set to Null if BlankMeasurement is False. Otherwise, automatically set to the value of Solvent in SamplesIn. If Solvent not specfied, set to Model[Sample, "Milli-Q water"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((Null | (ObjectP[{Model[Sample], Object[Sample]}] | _String)) | Automatic) | Null
Index Matches to: experiment samples
AbsorbanceBlankVolume
The volume of the blank that is transferred out and used for blank measurements prior to the Absorbance measurement. If AbsorbanceBlank is specified, AbsorbanceBlankVolume of Null indicates that the blanks are read inside their current containers. If AbsorbanceBlank is specified, AbsorbanceBlankVolume of a specific volume indicates the blanks are transferred to a container with the same model as the measurement container that holds the input cell samples or their aliquots for the Absorbance measurement.
Default Calculation: If BlankMeasurement is True, automatically set to the value of AssayVolume if that was specified, or maximum volume of the container otherwise.
Pattern Description: Greater than or equal to 1 microliter and less than or equal to 4000 microliters or Null.
Programmatic Pattern: ((Null | RangeP[1*Microliter, 4000*Microliter]) | Automatic) | Null
Index Matches to: experiment samples
NephelometryBlankMeasurement
Indicates if blank samples are prepared and measured prior to Nephelometry measurement. The scattered light attenuation of the blank samples are measured and background subtracted to account for any background signals.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
NephelometryBlank
The source used to generate a blank sample used to account for the background signal prior to the Nephelometry measurement of the cell samples of interest.
Default Calculation: Automatically set to Null if BlankMeasurement is False. Otherwise, automatically set to the value of Solvent in SamplesIn. If Solvent not specfied, set to Model[Sample, "Milli-Q water"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((Null | (ObjectP[{Model[Sample], Object[Sample]}] | _String)) | Automatic) | Null
Index Matches to: experiment samples
NephelometryBlankVolume
The volume of the blank that is transferred out and used for blank measurements prior to the Nephelometry measurement. If NephelometryBlank is specified, NephelometryBlankVolume of Null indicates that the blanks are read inside their current containers. If NephelometryBlank is specified, NephelometryBlankVolume of a specific volume indicates the blanks are transferred to a container with the same model as the measurement container that holds the input cell samples or their aliquots for the Nephelometry measurement.
Default Calculation: If BlankMeasurement is True, automatically set to the value of AssayVolume if that was specified, or maximum volume of the container otherwise.
Pattern Description: Greater than or equal to 1 microliter and less than or equal to 4000 microliters or Null.
Programmatic Pattern: ((Null | RangeP[1*Microliter, 4000*Microliter]) | Automatic) | Null
Index Matches to: experiment samples
Absorbance Measurement
AbsorbanceAcquisitionTemperature
Indicates the temperature the cell samples are held at during data acquisition within the absorbance instrument.
Default Calculation: Sets to Ambient if an instrument is capable of manipulating the temperature of the samples.
Pattern Description: Ambient or greater than or equal to 25 degrees Celsius and less than or equal to 45 degrees Celsius or Null.
Programmatic Pattern: ((RangeP[$AmbientTemperature, 45*Celsius] | Ambient) | Automatic) | Null
AbsorbanceEquilibrationTime
The length of time for which the cell samples are held at the requested temperature within the absorbance instrument before data acquisition.
Default Calculation: If an instrument is capable setting the temperature of the samples, sets to 0 second when Temperature is set to Ambient. Otherwise, it is set to 5 minutes.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 24 hours or Null.
Programmatic Pattern: ((Null | RangeP[0*Second, 24*Hour]) | Automatic) | Null
AbsorbanceTargetCarbonDioxideLevel
The target amount of carbon dioxide in the atmosphere in the plate reader chamber.
Default Calculation: Automatically set to 5% for mammalian cells, and Null otherwise.
Pattern Description: Greater than or equal to 0.1 percent and less than or equal to 20 percent or Null.
Programmatic Pattern: ((Null | RangeP[0.1*Percent, 20*Percent]) | Automatic) | Null
AbsorbanceTargetOxygenLevel
The target amount of oxygen in the atmosphere in the plate reader chamber. If specified, nitrogen gas is pumped into the chamber to force oxygen in ambient air out of the chamber until the desired level is reached.
Pattern Description: Greater than or equal to 0.1 percent and less than or equal to 20 percent or Null.
Programmatic Pattern: (Null | RangeP[0.1*Percent, 20*Percent]) | Null
AbsorbanceAtmosphereEquilibrationTime
The length of time for which the samples equilibrate at the requested oxygen and carbon dioxide level before being read.
Default Calculation: Automatically set to 5 Minute if TargetCarbonDioxideLevel or TargetOxygenLevel is specified. Otherwise, set to Null.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 24 hours or Null.
Programmatic Pattern: ((Null | RangeP[0*Second, 24*Hour]) | Automatic) | Null
NumberOfReadings
The number of times to acquire data from each input cell sample or its aliquot that is loaded into the instrument during the Absorbance measurement. Each data acquisition is performed on the same cell sample without reloading the instrument.
Default Calculation: If an instrument capable of adjusting NumberOfReadings is selected, resolves to 100. Otherwise resolves to Null
Pattern Description: Greater than or equal to 1 and less than or equal to 200 in increments of 1 or Null.
Programmatic Pattern: ((Null | RangeP[1, 200, 1]) | Automatic) | Null
AbsorbanceMethod
Indicates the type of container to be used to measure the absorbance of the input cell samples or its aliquot. PlateReader utilize an open well container that transverses light from top to bottom. Cuvette uses a square container with transparent sides to transverse light from the front to back at a fixed path length. Microfluidic uses small channels to load samples which are then gravity-driven towards chambers where light transverse from top to bottom and measured at a fixed path length.
Default Calculation: If any of the SamplesIn provided has a volume less than 500 Micro Liter, set to microfluidic. Otherwise, if there are less 8 samples, set to Cuvette. If none of options are true, set to PlateReader
Pattern Description: PlateReader or Cuvette or Null.
Programmatic Pattern: ((PlateReader | Cuvette) | Automatic) | Null
SpectralBandwidth
When using the Cuvette Method, indicates the physical size of the slit from which light passes out from the monochromator. The narrower the bandwidth, the greater the resolution in measurements.
Default Calculation: When using the Cuvette Method, automatically set 1.0 Nanometer. If using plate reader, set to Null.
Pattern Description: Greater than or equal to 0.5 nanometers and less than or equal to 5 nanometers or Null.
Programmatic Pattern: ((Null | RangeP[0.5*Nanometer, 5*Nanometer]) | Automatic) | Null
Wavelength
The specific wavelength which should be used to measure absorbance of the input cell samples or its aliquot.
Default Calculation: Automatically resolves to the shortest wavelength specified in the input samples' ExtinctionCoefficients field, and 260 Nanometer if that field is not populated.
Pattern Description: Greater than or equal to 200 nanometers and less than or equal to 1000 nanometers or Null.
Programmatic Pattern: (RangeP[200*Nanometer, 1000*Nanometer] | Automatic) | Null
Index Matches to: experiment samples
Nephelometry Measurement
NephelometryAcquisitionTemperature
Indicates the temperature the cell samples are held at during data acquisition within the nephelometer instrument.
Default Calculation: Automatically set to Ambient if Method->Solubility or the average of the IncubationTemperatures of the Model[Cell] Analytes, or 37 Celsius if that field is not informed if Method->CellCount.
Pattern Description: Ambient or greater than or equal to 25 degrees Celsius and less than or equal to 65 degrees Celsius or Null.
Programmatic Pattern: ((RangeP[$AmbientTemperature, 65*Celsius] | Ambient) | Automatic) | Null
NephelometryEquilibrationTime
The length of time for which the cell samples are held at the requested temperature within the nephelometer instrument before data acquisition.
Default Calculation: Automatically set to 0 second when Temperature is set to Ambient, or 5 minutes when Temperature is above Ambient.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 72 hours or Null.
Programmatic Pattern: (RangeP[0*Second, $MaxExperimentTime] | Automatic) | Null
NephelometryTargetCarbonDioxideLevel
The target amount of carbon dioxide in the atmosphere in the plate reader chamber.
Default Calculation: Automatically set to 5% for mammalian cells, and Null otherwise.
Pattern Description: Greater than or equal to 0.1 percent and less than or equal to 20 percent or Null.
Programmatic Pattern: ((Null | RangeP[0.1*Percent, 20*Percent]) | Automatic) | Null
NephelometryTargetOxygenLevel
The target amount of oxygen in the atmosphere in the plate reader chamber. If specified, nitrogen gas is pumped into the chamber to force oxygen in ambient air out of the chamber until the desired level is reached.
Pattern Description: Greater than or equal to 0.1 percent and less than or equal to 20 percent or Null.
Programmatic Pattern: (Null | RangeP[0.1*Percent, 20*Percent]) | Null
NephelometryAtmosphereEquilibrationTime
The length of time for which the samples equilibrate at the requested oxygen and carbon dioxide level before being read.
Default Calculation: Automatically set to 5 Minute if TargetCarbonDioxideLevel or TargetOxygenLevel is specified. Otherwise, set to Null.
Pattern Description: Greater than or equal to 0 seconds and less than or equal to 24 hours or Null.
Programmatic Pattern: ((Null | RangeP[0*Second, 24*Hour]) | Automatic) | Null
BeamAperture
The diameter of the opening allowing the source laser light to pass through to the cell sample. A larger BeamAperture allows more light to pass through to the sample, leading to a higher signal. A setting of 1.5 millimeters is recommended for all 384 and 96 well plates, and 2.5-3.5 millimeters for 48 or less well plates. For non-homogenous solutions, a higher BeamAperture is recommended, and for samples with a large meniscus effect, a smaller BeamAperture is recommended.
Pattern Description: Greater than or equal to 1.5 millimeters and less than or equal to 3.5 millimeters or Null.
Programmatic Pattern: (RangeP[1.5*Millimeter, 3.5*Millimeter] | Automatic) | Null
BeamIntensity
The percentage of the total amount of the laser source light passed through to reach the cell sample. For Solubility experiments, 80% is recommended, and for experiments with highly concentrated or highly turbid samples, such as those involving cells, a BeamIntensity of 10% is recommended.
Default Calculation: Automatically set to 80% if Method->Solubility, and 10% if Method->CellCount.
Pattern Description: Greater than or equal to 0 percent and less than or equal to 100 percent or Null.
Programmatic Pattern: (RangeP[0*Percent, 100*Percent] | Automatic) | Null
IntegrationTime
The amount of time that scattered light is measured. Increasing the IntegrationTime leads to higher signal and noise intensity.
Default Calculation: Automatically set to 1 second if no SamplingPattern is specified, or if SamplingPattern->Matrix. If SamplingPattern->Ring or Spiral, IntegrationTime is set based on the SamplingDistance.
Pattern Description: Greater than or equal to 20 milliseconds and less than or equal to 10 seconds or Null.
Programmatic Pattern: (RangeP[20*Millisecond, 10*Second] | Automatic) | Null