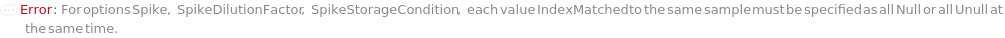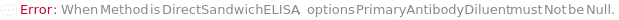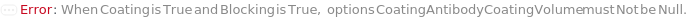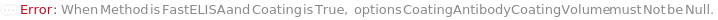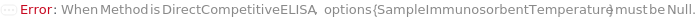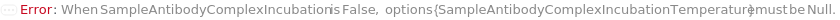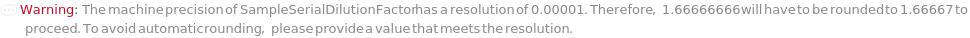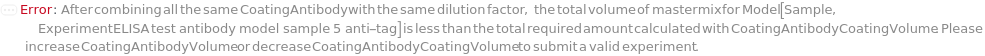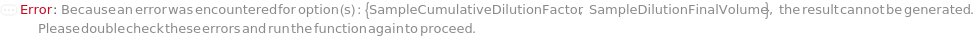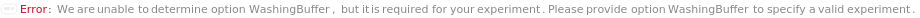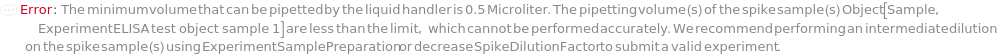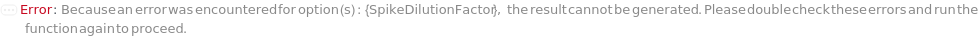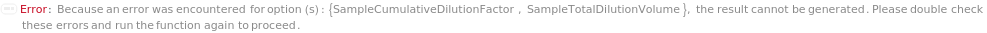General
Method
The format used for ELISA analysis in this experiment. Types include DirectELISA, IndirectELISA, DirectSandwichELISA, IndirectSandwichELISA, DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA. In a DirectELISA experiment, the sample is coated on the ELISAPlate, and then a primary antibody is used to detect the target antigen. In a DirectSandwichELISA experiment, a capture antibody is coated onto the ELISAPlate to pull down the target antigen from the sample, then a primary antibody is used to detect the target antigen. In a DirectCompetitiveELISA experiment, a reference antigen is used to coat the ELISAPlate, samples are incubated with the primary antibodies in the SampleAssemblyPlate to form Sample-Antibody-Complex, then, when the Sample-Antibody-Complex solution is loaded on the ELISAPlate, the remaining free primary antibody binds to the reference antigen. Compared with the direct methods where the primary antibody is conjugated with an enzyme (such as Horse Radish Peroxidase/HRP or Alkaline Phosphatase/AP) for detection, in all indirect methods a secondary antibody is, instead, conjugated with this enzyme, and an additional step of SecondaryAntibodyImmunosorbent is added to the corresponding direct methods. In a FastELISA experiment, a coating antibody against a tag is coated to the ELISAPlate, a capture antibody containing this tag and a primary antibody for antigen detection is incubated with the sample to form a CaptureAntibody-TargetAntigen-PrimaryAntibody complex in the SampleAssemblyPlate, and then this complex is loaded to the ELISAPlate and pulled down by the coating antibody to the surface of the plate. See Figure 3.1 and Figure 1.1-1.7 for more information about different ELISA methods.
Default Value: DirectELISA
Pattern Description: DirectELISA, IndirectELISA, DirectSandwichELISA, IndirectSandwichELISA, DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA.
Programmatic Pattern: ELISAMethodP
NumberOfReplicates
The number of times Samples will be repeated in parallel wells in an ELISA experiment. Replications are conducted at the same time, and if possible, in the same ELISAPlate. If set to Null, each Sample will be assayed once across all ELISA plates while each Standard or Blank is assayed once per ELISA plate. If set to a number, Samples are assayed multiple times cross all ELISA plates while each Standard or Blank is assayed once per ELISA plate. Up to three 96-well ELISA plates are allowed per experiment. See Figure 3.2 and 3.5 for more information about ELISAPlate(s) setup. Please use ExperimentELISAPreview to review the arrangement of Samples, Standards, and Blanks on the ELISA plates, and apply SamplesOutPlateAssignment, StandardDestinationWell, and BlankDestinationWell to adjust the well assignments as needed.
Pattern Description: Greater than or equal to 2 and less than or equal to 96 in increments of 1 or Null.
Programmatic Pattern: RangeP[2, 96, 1] | Null
Instrument
The instrument integrates a heater/cooler shaker, plate washer, and plate reader, and is used to dispense reagents into ELISA plates and transfer them between incubation and washing modules during the assay. See InstrumentTable section in the help file for details about integrated instruments and their operating limits.
Default Calculation: Automatically set to Model[Instrument, LiquidHandler, "Super STAR"].
Pattern Description: An object of type or subtype Model[Instrument, LiquidHandler] or Object[Instrument, LiquidHandler]
Programmatic Pattern: ObjectP[{Model[Instrument, LiquidHandler], Object[Instrument, LiquidHandler]}] | Automatic
DetectionMode
The type of detection method used to measure the signal generated from the enzyme substrate reaction to quantify the target analyte during the assay. Types include AbsorbanceIntensity for colorimetric signals, FluorescenceIntensity for fluorescent signals, and LuminescenceIntensity for chemiluminescent signals. See Figure 3.3 for more information.
Default Value: AbsorbanceIntensity
Pattern Description: AbsorbanceIntensity, FluorescenceIntensity, or LuminescenceIntensity.
Programmatic Pattern: AbsorbanceIntensity | FluorescenceIntensity | LuminescenceIntensity
NumberOfReadings
Number of redundant readings taken by the plate reader to determine a single averaged absorbance or fluorescence reading per well. This is currently available on "CLARIOstar" integrated to Model[Instrument, LiquidHandler, "Super STAR"]. This option is only applicable when DetectionMode is AbsorbanceIntensity or FluorescenceIntensity. For LuminescenceIntensity DetectionMode, please adjust IntegrationTime to increase signal-to-noise ratio.
Default Calculation: If the option DetectionMode is set to FluorescenceIntensity or AbsorbanceIntensity and the integrated plate reader instrument is capable of taking several readings per well, automatically set to 4 for AbsorbanceIntensity and 10 for FluorescenceIntensity.
Pattern Description: Greater than or equal to 1 and less than or equal to 200 or Null.
Programmatic Pattern: (RangeP[1, 200] | Automatic) | Null
Coating
Specifies whether ELISAPlate(s) require coating prior to the immunosorbent steps, indicating that the plates are not pre-coated. When pre-coated ELISA plates are not used, coating is required during this experiment. Coating is a procedure to immobilize coating analytes (usually antigens and antibodies) to the surface of the assay plate(s) through non-specific adsorption. The ELISAPlate(s) are coated with samples for DirectELISA or IndirectELISA, CaptureAntibodies for DirectSandwichELISA or IndirectSandwichELISA, ReferenceAntigens for DirectCompetitiveELISA or IndirectCompetitiveELISA, and CoatingAntibodies for FastELISA. See Figure 3.1 for more information about ELISAPlate configurations across different methods.
Default Calculation: If ELISAPlate is specified and contains any contents, automatically set to False. If the specified ELISAPlate model has the field Coating populated with analytes, automatically set to False. Otherwise set to True.
Pattern Description: True or False.
Programmatic Pattern: BooleanP | Automatic
Blocking
Indicates whether BlockingBuffer is added to the ELISAPlate(s) to saturate unoccupied binding sites and prevent nonspecific binding of assay components.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
SampleAssemblyPlate
The desired plate model used for preparing samples, standards, and blanks, including performing dilutions, spiking and mixing with other reagents (such as PreImmunosorbentLigand, PrimaryAntibody, and CaptureAntibody). This supports achieving the desired sample concentrations and, when applicable, the formation and incubation of sample antibody or sample ligand complexes prior to transfer to ELISAPlate(s). The SampleAssemblyPlate(s) will be discarded at the end of the experiment.
Default Calculation: Automatically set SampleAssemblyPlate to Model[Container, Plate, "96-well 2mL Deep Well Plate"] if the method is DirectELISA or IndirectELISA and at least one sample or standard requires serial or linear dilution, or if the method is DirectSandwichELISA or IndirectSandwichELISA and either at least one sample or standard requires serial or linear dilution or a PreImmunosorbentLigand is specified, or if the method is DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA.
Pattern Description: An object of type or subtype Model[Container, Plate] or Null.
Programmatic Pattern: (ObjectP[Model[Container, Plate]] | Automatic) | Null
ELISAPlate
The 96-well flat-bottom plate that serves as the solid-phase support for the enzyme-linked immunosorbent assay, where samples, standards, and blanks are loaded and the immunosorbent reactions take place. The same plate serves as the platform for signal measurement. Please use ExperimentELISAPreview to check the arrangement of Samples, Standards, and Blanks on the ELISAPlate.
Default Calculation: If samples are pre-coated to a plate, ELISAPlate is automatically set to the corresponding pre-coated plate. Otherwise, ELISAPlate is automatically selected based on DetectionMode. When DetectionMode is AbsorbanceIntensity, ELISAPlate is automatically set to Model[Container, Plate, "96-well Polystyrene Flat-Bottom Plate, Clear"], which is optimized for absorbance measurements. When DetectionMode is FluorescenceIntensity, ELISAPlate is automatically set to Model[Container, Plate, "96-well Black Wall Greiner Plate"], which minimizes cross-talk and reduces background fluorescence. When DetectionMode is LuminescenceIntensity, ELISAPlate is automatically set to Model[Container, Plate, "96-well plate, High Binding, White Opaque"], which minimizes cross-talk and enhances luminescence signal reflection.
Pattern Description: An object of type or subtype Model[Container, Plate] or Object[Container, Plate] or a prepared sample.
Programmatic Pattern: (ObjectP[{Model[Container, Plate], Object[Container, Plate]}] | _String) | Automatic
SecondaryELISAPlate
The second assay plate that serves as the solid-phase support for the enzyme-linked immunosorbent assay, where samples, standards, and blanks are loaded and the immunosorbent reactions take place, needed if the total number of Sample (including spiked samples and dilutions) times NumberOfReplicates plus standards, blanks, and moating samples exceeds 96. See Figure 3.2 for more information about ELISAPlate(s) setup with different dilution, replication and moating options. Please use ExperimentELISAPreview to check the arrangement of Samples, Standards, and Blanks on the SecondaryELISAPlate.
Default Calculation: If samples are pre-coated to more than one plate, SecondaryELISAPlate is automatically set to the second corresponding pre-coated plate. If the total number of Sample (including spiked samples and dilutions) times NumberOfReplicates plus standards, blanks, and moating samples exceeds 96, SecondaryELISAPlate is automatically set to the same container model as ELISAPlate.
Pattern Description: An object of type or subtype Model[Container, Plate] or Object[Container, Plate] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Container, Plate], Object[Container, Plate]}] | _String) | Automatic) | Null
TertiaryELISAPlate
The third assay plate that serves as the solid-phase support for the enzyme-linked immunosorbent assay, where samples, standards, and blanks are loaded and the immunosorbent reactions take place, needed if the total number of Sample (including spiked samples and dilutions) times NumberOfReplicates plus standards, blanks, and moating samples exceeds 192. See Figure 3.2 for more information about ELISAPlate(s) setup with different different dilution, replication and moating options. Please use ExperimentELISAPreview to check the arrangement of Samples, Standards, and Blanks on the TertiaryELISAPlate.
Default Calculation: If samples are pre-coated to more than two plates, TertiaryELISAPlate is automatically set to the third corresponding pre-coated plate. If the total number of Sample (including spiked samples and dilutions) times NumberOfReplicates plus standards, blanks, and moating samples exceeds 192, TertiaryELISAPlate is automatically set to the same model as ELISAPlate.
Pattern Description: An object of type or subtype Model[Container, Plate] or Object[Container, Plate] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Container, Plate], Object[Container, Plate]}] | _String) | Automatic) | Null
TargetAntigen
The analyte molecule (e.g., peptide, protein, or hormone) detected and quantified in the samples by antibodies in this assay. This option is used to automatically set antibody options (such as PrimaryAntibody, CaptureAntibody, CoatingAntibody) and the corresponding experiment conditions of Standards and Blanks.
Default Calculation: Automatically set to the first Model[Molecule, Protein] in the Analytes field of the input sample, or, if not populated, to the first Model[Molecule, Protein] in the Composition field of the input sample, or if none exist, set to Null.
Pattern Description: An object of type or subtype Model[Molecule] or Null.
Programmatic Pattern: (ObjectP[Model[Molecule]] | Automatic) | Null
Index Matches to: experiment samples
Sample Assembly
Spike
The sample with a known concentration of analyte. Spike is to be mixed with the undiluted input sample based on SpikeDilutionFactor. The purpose of spiking is to perform a spike-and-recovery assessment to determine whether the ELISA can accurately test the concentration of analyte within the context of the sample. If the recovery observed for the spiked sample is identical to the known concentration, the sample is considered not to interfere with the assay. For example, if molecules in the sample inhibits the binding between the antibody and TargetAntigen, the results from the ELISA becomes inaccurate. In a Spike-and-Recovery experiment, ELISA is performed on the Sample alone (also called Neat Sample) and the Spiked Sample at the same time, where the Spike concentration in the sample can be measured. This measured Spike concentration can be then compared with the known Spike concentration. Spiked sample can be further serial diluted the same way as the neat sample to perform linearity-of-dilution assessment. The SampleAssemblyPlate used for sample spiking and dilution will be discarded after the experiment.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: (ObjectP[{Model[Sample], Object[Sample]}] | _String) | Null
Index Matches to: experiment samples
SpikeDilutionFactor
The factor by which the Spike is mixed with the input sample before further dilution is performed, calculated as the total final volume consisting of both the Spike and the input sample divided by the volume of the Spike. For example, a ten-fold dilution is corresponding to SpikeDilutionFactor at 10. SpikeDilutionFactor must be specified whenever Spike is specified.
Default Calculation: Automatically set to 10 if Spike is not Null and input sample is in liquid state.
Pattern Description: Greater than 1 or Null.
Programmatic Pattern: (GreaterP[1] | Automatic) | Null
Index Matches to: experiment samples
SampleDilutionType
Indicates whether the sample is diluted, and if so, specifies the dilution type (linear or serial). Linear dilution represents a single stage dilution of the TargetAntigen in the sample by specified SampleDilutionTransferVolume and SampleDilutionDiluentVolume. For example, a two-fold, five-fold and ten-fold linear dilution takes 500 Microliters, 200 Microliters and 100 Microliters of the input sample and generates three final diluted samples at 1 Milliliter (set by SampleTotalDilutionVolume at 1 Milliliter, SampleCumulativeDilutionFactor at {2, 5, 10} and SampleNumberOfDilutions at 3). Serial dilution represents a stepwise dilution of TargetAntigen in the sample resulting in multiple samples and a geometric progression of the concentration. The first source sample is the original sample provided. For example, a sample with initial TargetAntigen concentration of 100ng/ul, if diluted 3 times (set by SampleNumberOfDilutions at 3) and two-fold dilution at each stage (set by SampleSerialDilutionFactor at 2), should yield ELISA measurements of 50ng/ul, 25ng/ul, and 12.5ng/ul when SampleDilutionStrategy is Series or just 12.5ng/ul when SampleDilutionStrategy is Endpoint. See Figure 3.4 for more information about Linear and Serial dilution workflow.
Default Calculation: Automatically set to None if Method is not DirectELISA or IndirectELISA and Coating is not False.
Pattern Description: {Serial, Linear} or None or Null.
Programmatic Pattern: ((DilutionTypeP | None) | Automatic) | Null
Index Matches to: experiment samples
SampleDilutionStrategy
Indicates if only the final diluted sample (Endpoint) or all diluted samples (Series) produced by serial dilution are quantified in ELISA. This option is only applicable when SampleDilutionType is Serial. See Figure 3.2 for more information about ELISAPlate(s) setup with different SampleDilutionStrategies.
Default Calculation: Automatically set to Series if SampleDilutionType is Serial.
Pattern Description: Series or Endpoint or Null.
Programmatic Pattern: ((Null | DilutionStrategyP) | Automatic) | Null
Index Matches to: experiment samples
SampleNumberOfDilutions
The number of diluted samples to prepare. When SampleDilutionStrategy is Linear, all prepared diluted samples are directly diluted from the undiluted Sample but can be different in SampleCumulativeDilutionFactor. When SampleDilutionStrategy is Serial, the dilution is performed stepwise, producing samples with progressively decreasing concentrations across all dilution levels. See Figure 3.2 to see examples with different SampleDilutionStrategies and SampleNumberOfDilutions.
Default Calculation: Automatically set to the length of SampleCumulativeDilutionFactor, or SampleSerialDilutionFactor if provided, or 1 if not provided. See Figure 3.4 for more information.
Pattern Description: Greater than or equal to 1 and less than or equal to 15 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 15, 1] | Automatic) | Null
Index Matches to: experiment samples
SampleCumulativeDilutionFactor
The factor by which the concentration of the TargetAntigen in the original sample is reduced during the dilution. The length of this list must match the corresponding value in SampleNumberOfDilutions. In the example in the option SampleDilutionType when it is set to Serial, two-fold dilution for 3 times is corresponding to SampleCumulativeDilutionFactor at {2,4,8}. When SampleDilutionType is Linear, each dilution is made independently from the original sample. In the example in the option SampleDilutionType when it is set to Linear, a two-fold, five-fold and ten-fold linear dilution is corresponding to SampleCumulativeDilutionFactor at {2,5,10}.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 1 and less than or equal to 100000000000000000000000 or Null.
Programmatic Pattern: (RangeP[1, 10^23] | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SampleSerialDilutionFactor
The factor by which the concentration of the TargetAntigen in the resulting sample of the previous dilution step is reduced. For example, if the SampleCumulativeDilutionFactor is equal to {2,4,8}, SampleSerialDilutionFactor is 2 or {2,2,2}. Because the dilution curve does not intrinsically include the original (undiluted) sample, the first dilution factor must be set to 1 if the original sample is to be measured as well. Using the same example described in SampleDilutionType, to obtain ELISA measurements of 100 ng/ul, 50 ng/uL, 25 ng/ul, and 12.5 ng/ul, SampleSerialDilutionFactor should be defined as {1, 2, 2, 2}, and SampleNumberOfDilutions should be set to 4. This option is only applicable when SampleDilutionType is Serial.
Default Calculation: Automatically set based on the systems of equations specified under the SampleDilutionType option. If no sample dilution options are specified, automatically set to 10 when SampleDilutionStrategy is Serial.
Pattern Description: Greater than or equal to 1 and less than or equal to 100000000000000000000000 or Null.
Programmatic Pattern: ((Null | RangeP[1, 10^23]) | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SampleDilutionTransferVolume
The amount of sample that is diluted with SampleDiluent when SampleDilutionType is set to Linear, or the amount of sample transferred from the resulting sample of one round of the dilution series to the next sample in the series when SampleDilutionType is set to Serial. When a Spike is added to and mixed with the input sample, its volume contributes to the total SampleDilutionTransferVolume used for dilution calculations.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SampleDilutionDiluentVolume
The amount of SampleDiluent added to dilute the sample when SampleDilutionType is set to Linear, or the amount of SampleDiluent added to dilute the sample at each stage of the dilution when SampleDilutionType is set to Serial.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: ((Null | RangeP[0*Milliliter, $MaxTransferVolume]) | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SampleTotalDilutionVolume
The combined volume of the input sample and SampleDiluent. If SampleDilutionType is set to Serial, this is the volume of the resulting sample before SampleDilutionTransferVolume is removed for use in the next dilution in the series. If SampleDilutionType is set to Linear, this is the sum of SampleDilutionDiluentVolume and SampleDilutionTransferVolume.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SampleDilutionFinalVolume
The volume of the resulting diluted sample after SampleDilutionTransferVolume is removed for use in the next dilution in the series if SampleDilutionType is set to Serial. This option is only applicable when SampleDilutionType is Serial.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
Nested Index Matches to: experiment samples
SamplesOutPlateAssignment
The well and plate locations assigned for each output sample on the ELISA plates, corresponding to the given input sample. This includes the neat sample, all diluted series, and spiked sample, if applicable, and spiked dilutions, in that order. SamplesOutPlateAssignment can be customized. When configuring SamplesOutPlateAssignment, set IndexMatch to Variable and ensure that the number of {DestinationWell, Plate} entries exactly matches the total number of SamplesOut. Any replicates specified by NumberOfReplicates must also be accounted for in the assignment. Please use ExperimentELISAPreview to verify the arrangement of Samples on the ELISA plates. If SamplesOutPlateAssignment is not specified, default well assignments will be automatically generated in a column-wise fashion. See Figure 3.5 for examples of the default assignment.
Default Calculation: If Method is DirectELISA or IndirectELISA and Coating is False, automatically set to the well and plate position of the input sample. Otherwise, filling the ELISA plates in the order of Blank and Standard first, then starting the well assignment for samples from the first available well, and continue columnwise. Replications are placed at the same time in sequential positions if possible.
Pattern Description: List of one or more {DestinationWell, Plate} entries or Null.
Programmatic Pattern: ({{Alternatives @@ Flatten[AllWells[NumberOfWells -> 96]], ELISAPlate | SecondaryELISAPlate | TertiaryELISAPlate}..} | Automatic) | Null
Index Matches to: experiment samples
SampleDiluent
The buffer used to dilute samples (both neat samples and spiked samples) to appropriate concentrations for the assay. It provides a controlled chemical environment that maintains protein stability, preserves antigen antibody binding conditions, and minimizes nonspecific interactions. This option is only applicable when SampleDilutionType is not None.
Default Calculation: If SampleDilutionType is not None or Null for all input samples, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"] when Method is set to DirectELISA or IndirectELISA, otherwise set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Sample Antibody Complex Incubation
SampleVolume
The volume of prepared sample(s) (both neat sample, spiked sample, and all diluted samples if applicable) that are dispensed into the SampleAssemblyPlate(s) in order to form Sample-Antibody complex or Sample-Ligand complex.
Default Calculation: If Method is FastELISA, DirectCompetitiveELISA, or IndirectCompetitiveELISA, automatically set to 100 Microliters. If Method is DirectSandwichELISA or IndirectSandwichELISA and PreImmunosorbentLigand is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
SampleAntibodyComplexIncubation
Specifies whether the pre-mixed samples and antibodies in the SampleAssemblyPlate(s) undergo an incubation step to facilitate the formation of the sample antibody complex prior to transferring into the ELISAPlate(s). In DirectCompetitiveELISA and IndirectCompetitiveELISA, PrimaryAntibodies are premixed with the input samples, standards, and blanks. In FastELISA, PrimaryAntibodies and CaptureAntibodies are premixed with the input samples, standards, and blanks. If set to False, the prepared sample-antibody complex will be dispensed to ELISAPlate(s) immediately. This option is only applicable when Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA.
Default Calculation: Automatically set to True if Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA and neither SampleAntibodyComplexIncubationTime nor SampleAntibodyComplexIncubationTemperature is Null.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SampleAntibodyComplexIncubationTime
The incubation duration of pre-mixed samples and antibodies during SampleAntibodyComplexIncubation. Incubation time depends on the antigen-antibody affinity, concentration of the samples and antibodies, and temperature.
Default Calculation: If SampleAntibodyComplexIncubation is True, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or Null.
Programmatic Pattern: (RangeP[0*Minute, 24*Hour] | Automatic) | Null
SampleAntibodyComplexIncubationTemperature
The temperature at which the pre-mixed samples and antibodies are incubated during SampleAntibodyComplexIncubation.
Default Calculation: If SampleAntibodyComplexIncubation is True, automatically set to Ambient.
Pattern Description: Ambient or greater than or equal to 4 degrees Celsius and less than or equal to 50 degrees Celsius or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
SampleAntibodyComplexIncubationMixRate
The speed at which the SampleAssemblyPlate(s) containing the pre-mixed samples and antibodies is shaken (orbitally, at a radius of 2 mm) during SampleAntibodyComplexIncubation. Gentle orbital shaking helps maintain homogeneity and speeds up binding. Shaking too vigorously, however, can cause splashing or evaporation, leading to inconsistent results.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
Coating
SampleCoatingVolume
The amount of prepared sample(s) (both neat sample, spiked sample, and all diluted samples if applicable) that are dispensed into the ELISAPlate(s), in order for the Sample to be adsorbed to the surface of the well.
Default Calculation: If Method is DirectELISA or IndirectELISA and Coating is True, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
CoatingAntibodyCoatingVolume
The amount of CoatingAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the CoatingAntibody to be adsorbed to the surface of the well.
Default Calculation: If CoatingAntibody is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
CaptureAntibodyCoatingVolume
The amount of CaptureAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the CaptureAntibody to be adsorbed to the surface of the well. This option is only applicable when Method is DirectSandwichELISA or IndirectSandwichELISA.
Default Calculation: If CaptureAntibody is specified and Method is DirectSandwichELISA or IndirectSandwichELISA, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
ReferenceAntigenCoatingVolume
The amount of diluted ReferenceAntigen (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the ReferenceAntigen to be adsorbed to the surface of the well.
Default Calculation: If ReferenceAntigen is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
CoatingTemperature
The temperature at which the ELISAPlate(s) are kept during coating, in order for the coating analytes to be adsorbed to the surface of the assay plate(s).
Default Calculation: If Coating is set to True, automatically set to 37 degrees Celsius.
Pattern Description: Ambient or greater than or equal to 4 degrees Celsius and less than or equal to 50 degrees Celsius or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
CoatingTime
The duration for coating step. The coated ELISAPlate(s) will be kept at 4 degree Celsius after the CoatingTime is reached or immediately blocked. Coating time depends on the properties of the assay plate(s), hydrophobicity, stability, and concentration of the coating analytes, and temperature.
Default Calculation: If Coating is set to True, automatically set to 1 hour when CoatingTemperature is Ambient or above, set to 16 hours when CoatingTemperature is below 8 degrees celsius, or set to 8 hours when CoatingTemperature is in between.
Pattern Description: Greater than or equal to 0 hours and less than or equal to 20 hours or Null.
Programmatic Pattern: (RangeP[0*Hour, 20*Hour] | Automatic) | Null
CoatingWashing
Indicates whether a final washing step is performed before blocking to remove unbound coating analytes from either pre-coated ELISAPlate(s) or ELISAPlate(s) coated during the experiment. CoatingWashing is usually recommended when Coating is set to True, or when Coating is set to False but Blocking is set to True. Pre-coated plate(s) are subjected to this washing step even though no coating incubation is performed.
Default Calculation: If Coating is set to True, automatically set to True. If Coating is set to False but Blocking is set to True, automatically set to True. Otherwise, set to False.
Pattern Description: True or False.
Programmatic Pattern: BooleanP | Automatic
CoatingWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off unbound coating analytes from the ELISAPlate(s). When Coating is set to False but Blocking is set to True, the pre-coated plate(s) are subjected to the wash step during coating, despite the absence of a coating incubation step.
Default Calculation: If CoatingWashing is True, automatically set to 250 Microliters per well.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
CoatingNumberOfWashes
The number of washes performed to rinse off unbound coating analytes. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the ELISAPlate(s).
Default Calculation: If CoatingWashing, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
MoatSize
Indicates the number of concentric perimeters of wells which should be should be filled with MoatingBuffer in order to reduce evaporation from the assay samples during the coating incubation. For example, when MoatSize is set to 1, wells A1-H1, A1-A12, H1-H12, A12-H12 are filled with MoatingBuffer. This option is only applicable when Method is DirectELISA or IndirectELISA and Coating is set to True. See Figure 1.1-1.2 and 3.6 for more information about sample preparation with moating for DirectELISA and IndirectELISA procedures.
Pattern Description: Greater than or equal to 1 and less than or equal to 3 in increments of 1 or Null.
Programmatic Pattern: RangeP[1, 3, 1] | Null
MoatVolume
Indicates the volume which should be added to each moat well.
Default Calculation: Automatically set to 100 Microliters if MoatSize is specified.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
MoatingBuffer
Indicates the buffer which should be used to fill each moat well.
Default Calculation: Automatically set to Model[Sample, "ELISA Blocker Blocking Buffer"] if MoatSize is specified.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Immunosorbent Step
SampleImmunosorbentVolume
The volume of the prepared sample(s) (both neat sample, spiked sample, and all diluted samples if applicable) to be loaded on the ELISAPlate(s) for the TargetAntigen to bind to the CaptureAntibody in DirectSandwichELISA and IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
PrimaryAntibodyImmunosorbentVolume
The volume of the PrimaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for immunosorbent step when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA, set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
SecondaryAntibodyImmunosorbentVolume
The volume of the SecondaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for the immunosorbent step.
Default Calculation: If SecondaryAntibody is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
SampleAntibodyComplexImmunosorbentVolume
The volume of the sample-antibody complex to be loaded on the ELISAPlate(s). In DirectCompetitiveELISA and IndirectCompetitiveELISA, this step enables the free PrimaryAntibody to bind to the ReferenceAntigen coated on the ELISAPlate(s). In FastELISA, this step enables the PrimaryAntibody-TargetAntigen-CaptureAntibody complex to bind to the CoatingAntibody on the ELISAPlate(s).
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
SampleAntibodyComplexImmunosorbentTime
The duration for which the sample-antibody complex is allowed to adsorb onto the ELISAPlate(s).
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or Null.
Programmatic Pattern: (RangeP[0*Minute, 24*Hour] | Automatic) | Null
SampleAntibodyComplexImmunosorbentTemperature
The temperature at which the ELISAPlate(s) are kept during SampleAntibodyComplexImmunosorbent incubation.
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
SampleAntibodyComplexImmunosorbentMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during SampleAntibodyComplexImmunosorbent incubation.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
SampleAntibodyComplexImmunosorbentWashing
Indicates if a final washing step is performed at the end of SampleAntibodyComplexImmunosorbent incubation to wash off unbound materials. Performing this washing step is generally recommended. However, some specialized commercial pre-blocked plates are designed with detection chemistry specific to the bound complex, in which case the washing step may be optional.
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SampleAntibodyComplexImmunosorbentWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off the unbound materials from ELISAPlate(s) after SampleAntibodyComplexImmunosorbent incubation.
Default Calculation: If SampleAntibodyComplexImmunosorbentWashing is set to True, automatically set to 250 Microliter.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
SampleAntibodyComplexImmunosorbentNumberOfWashes
The number of rinses performed after SampleAntibodyComplexImmunosorbent step. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the ELISAPlate(s).
Default Calculation: If SampleAntibodyComplexImmunosorbentWashing is set to True, the option is automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
SampleImmunosorbentTime
The duration for which the samples are allowed to adsorb onto the assay plate(s) in DirectSandwichELISA and IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or Null.
Programmatic Pattern: (RangeP[0*Minute, 24*Hour] | Automatic) | Null
SampleImmunosorbentTemperature
The temperature at which the ELISAPlate(s) are kept during SampleImmunosorbent incubation.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
SampleImmunosorbentMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during SampleImmunosorbent incubation.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
SampleImmunosorbentWashing
Indicates if a final washing step is performed at the end of SampleImmunosorbent incubation to wash off unbound samples. Performing this washing step is generally recommended. However, some specialized commercial pre-blocked plates are designed with detection chemistry specific to the bound complex, in which case the washing step may be optional.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SampleImmunosorbentWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off the unbound Samples after SampleImmunosorbent incubation.
Default Calculation: If SampleImmunosorbentWashing is set to True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
SampleImmunosorbentNumberOfWashes
The number of rinses performed to rinse off the unbound Samples from ELISAPlate(s) after SampleImmunosorbent incubation. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the assay plate(s).
Default Calculation: If SampleImmunosorbentWashing is set to True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
PrimaryAntibodyImmunosorbentTime
The duration for which the PrimaryAntibodies are allowed to adsorb onto the ELISAPlate(s) for DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or Null.
Programmatic Pattern: (RangeP[0*Minute, 24*Hour] | Automatic) | Null
PrimaryAntibodyImmunosorbentTemperature
The temperature at which the assay plate(s) are kept during PrimaryAntibodyImmunosorbent incubation.
Default Calculation: If the Method is set to DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
PrimaryAntibodyImmunosorbentMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during PrimaryAntibodyImmunosorbent incubation.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
PrimaryAntibodyImmunosorbentWashing
Indicates if a final washing step is performed at the end of PrimaryAntibodyImmunosorbent incubation to wash off unbound PrimaryAntibody. Performing this washing step is generally recommended. However, some specialized commercial pre-blocked plates are designed with detection chemistry specific to the bound complex, in which case the washing step may be optional.
Default Calculation: If Method is set to DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
PrimaryAntibodyImmunosorbentWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off the unbound PrimaryAntibody after PrimaryAntibodyImmunosorbent incubation.
Default Calculation: If PrimaryAntibodyImmunosorbentWashing is True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
PrimaryAntibodyImmunosorbentNumberOfWashes
The number of rinses performed to rinse off the unbound PrimaryAntibody from assay plate(s) after PrimaryAntibodyImmunosorbent incubation. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the ELISAPlate(s).
Default Calculation: If PrimaryAntibodyImmunosorbentWashing is True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
SecondaryAntibodyImmunosorbentTime
The duration for which the SecondaryAntibodies are allowed to adsorb onto the ELISAPlate(s) for IndirectELISA, IndirectSandwichELISA, and IndirectCompetitiveELISA.
Default Calculation: If the Method is set to IndirectELISA, IndirectSandwichELISA, and IndirectCompetitiveELISA, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or Null.
Programmatic Pattern: (RangeP[0*Minute, 24*Hour] | Automatic) | Null
SecondaryAntibodyImmunosorbentTemperature
The temperature at which the ELISAPlate(s) are kept during SecondaryAntibodyImmunosorbent incubation.
Default Calculation: If the Method is set to IndirectELISA, IndirectSandwichELISA, and IndirectCompetitiveELISA, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
SecondaryAntibodyImmunosorbentMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during SecondaryAntibodyImmunosorbent incubation.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
SecondaryAntibodyImmunosorbentWashing
Indicates if a final washing step is performed at the end of SecondaryAntibodyImmunosorbent incubation to wash off unbound SecondaryAntibody. Performing this washing step is generally recommended. However, some specialized commercial pre-blocked plates are designed with detection chemistry specific to the bound complex, in which case the washing step may be optional.
Default Calculation: If Method is set to IndirectELISA, IndirectSandwichELISA, and IndirectCompetitiveELISA, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SecondaryAntibodyImmunosorbentWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off the unbound Secondary antibody after SecondaryAntibodyImmunosorbent incubation.
Default Calculation: If SecondaryAntibodyImmunosorbentWashing is True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
SecondaryAntibodyImmunosorbentNumberOfWashes
The number of rinses performed to rinse off the unbound SecondaryAntibody from ELISAPlate(s) after SecondaryAntibodyImmunosorbent incubation. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the assay plate(s).
Default Calculation: If SecondaryAntibodyImmunosorbentWashing is True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
FastELISA Reagent Preparation
CoatingAntibody
The antibody to be immobilized on the surface of the ELISA plate(s) to capture the TargetAntigen during the assay. This option is only applicable when Method is FastELISA.
Default Calculation: If Method is FastELISA and Coating is True, automatically set to an antibody against the affinity label tag which is conjugated to the CaptureAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: experiment samples
CoatingAntibodyDilutionFactor
The factor by which the CoatingAntibody concentration is reduced by CoatingAntibodyDiluent, calculated as the total final volume consisting of both the CoatingAntibody and CoatingAntibodyDiluent divided by the volume of the CoatingAntibody. When CoatingAntibodyDilutionFactor is 1, no dilution is performed. When preparing the coating antibody master mix in CoatingAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same CoatingAntibody and CoatingAntibodyDilutionFactor. For example, if CoatingAntibodyDilutionFactor is 100 and CoatingAntibodyVolume is 5 Microliters, the coating antibody master mix is prepared by combining 5 Microliters of undiluted CoatingAntibody with 495 Microliters of CoatingAntibodyDiluent, assuming no duplicate dilutions are made. If two samples share the same CoatingAntibody and its dilution factor, the master mix is prepared by mixing 10 Microliters of undiluted CoatingAntibody with 990 Microliters of CoatingAntibodyDiluent. This option is only applicable when Method is FastELISA.
Default Calculation: Automatically set to 1000 when CoatingAntibody is specified.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: experiment samples
CoatingAntibodyVolume
The volume of undiluted CoatingAntibody directly added into the CoatingAntibodyDilutionContainers to mix with CoatingAntibodyDiluent corresponding to the input sample. This option is only applicable when CoatingAntibodyDilutionFactor is not Null or 1 and CoatingAntibody is specified.
Default Calculation: If CoatingAntibody is specified and CoatingAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*CoatingAntibodyCoatingVolume/CoatingAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
CoatingAntibodyDiluent
The buffer used to dilute CoatingAntibody as well as StandardCoatingAntibody and BlankCoatingAntibody to appropriate concentrations for the assay before plate coating. It provides conditions that support efficient antibody adsorption to the plate surface while preserving antibody stability.
Default Calculation: If coating antibodies and their dilutionFactors for samples, standards and blanks are specified, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
FastELISA/SandwichELISA Reagent Preparation
CaptureAntibody
The antibody that is used to pull down the TargetAntigen from sample solution to the surface of the ELISAPlate(s) in DirectSandwichELISA, IndirectSandwichELISA, and FastELISA methods.
Default Calculation: If Method is FastELISA, automatically set to an antibody containing an affinity tag and against the TargetAntigen. If Method is DirectSandwichELISA and Coating is True, automatically set to an un-tagged antibody against TargetAntigen. If Method is IndirectSandwichELISA and Coating is True, automatically set to an untagged antibody against TargetAntigen but not a target for the SecondaryAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: experiment samples
CaptureAntibodyVolume
The volume of undiluted CaptureAntibody directly added into the corresponding well of the CaptureAntibodyDilutionContainers to mix with CaptureAntibodyDiluent when Method is DirectSandwichELISA or IndirectSandwichELISA, or the volume of undiluted CaptureAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to form Sample-Antibody complex when Method is FastELISA.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and CaptureAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*CaptureAntibodyCoatingVolume/CaptureAntibodyDilutionFactor. If Method is FastELISA, automatically set to 2 Microliters.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
SandwichELISA Reagent Preparation
CaptureAntibodyDilutionFactor
The factory by which the CaptureAntibody concentration is reduced by CaptureAntibodyDiluent, calculated as the total final volume consisting of both the CaptureAntibody and CaptureAntibodyDiluent divided by the volume of the CaptureAntibody. When CaptureAntibodyDilutionFactor is 1, no dilution is performed. When preparing the capture antibody master mix in CaptureAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same CaptureAntibody and CaptureAntibodyDilutionFactor. For example, if CaptureAntibodyDilutionFactor is 100 and CaptureAntibodyVolume is 5 Microliters, the capture antibody master mix is prepared by combining 5 Microliters of undiluted CaptureAntibody with 495 Microliters of CaptureAntibodyDiluent, assuming no duplicate dilutions are made. If two samples share the same CaptureAntibody and its dilution factor, the master mix is prepared by mixing 10 Microliters of undiluted CaptureAntibody with 990 Microliters of CaptureAntibodyDiluent. This option is only applicable for DirectSandwichELISA and IndirectSandwichELISA methods. For FastELISA method, CaptureAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: For Method is DirectSandwichELISA and IndirectSandwichELISA, automatically set to 1000 if CaptureAntibody is specified.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: experiment samples
CaptureAntibodyDiluent
The buffer used to dilute CaptureAntibody as well as StandardCaptureAntibody and BlankCaptureAntibody to appropriate concentrations for the assay before plate coating when Method is DirectSandwichELISA or IndirectSandwichELISA.
Default Calculation: If capture antibodies and their dilutionFactors for samples, standards and blanks are specified, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
CompetitiveELISA Reagent Preparation
ReferenceAntigen
The antigen that competes with TargetAntigen in the sample for the binding of the PrimaryAntibody. It is also referred to as inhibitor antigen. This option is only applicable when Method is DirectCompetitiveELISA or IndirectCompetitiveELISA.
Default Calculation: If Method is set to DirectCompetitiveELISA or IndirectCompetitiveELISA, automatically set to a sample containing known amount of TargetAntigen if Coating is True.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: experiment samples
ReferenceAntigenDilutionFactor
The factor by which the ReferenceAntigen concentration is reduced by ReferenceAntigenDiluent, calculated as the total final volume consisting of both the ReferenceAntigen and ReferenceAntigenDiluent divided by the volume of the ReferenceAntigen. When ReferenceAntigenDilutionFactor is 1, no dilution is performed. When preparing the reference antigen master mix in ReferenceAntigenDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same ReferenceAntigen and ReferenceAntigenDilutionFactor. For example, if ReferenceAntigenDilutionFactor is 100 and ReferenceAntigenVolume is 5 Microliters, the reference antigen master mix is prepared by combining 5 Microliters of undiluted ReferenceAntigen with 495 Microliters of ReferenceAntigenDiluent, assuming no duplicate dilutions are made. If two samples share the same ReferenceAntigen and its dilution factor, the master mix is prepared by mixing 10 Microliters of undiluted ReferenceAntigen with 990 Microliters of ReferenceAntigenDiluent. This option is only applicable for DirectCompetitiveELISA or IndirectCompetitiveELISA methods.
Default Calculation: If ReferenceAntigen is specified, automatically set to 1000.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: experiment samples
ReferenceAntigenVolume
The volume of undiluted ReferenceAntigen directly added into the ReferenceAntigenDilutionContainers to mix with ReferenceAntigenDiluent corresponding to the input sample. This option is only applicable when ReferenceAntigenDilutionFactor is not Null or 1 and ReferenceAntigen is specified.
Default Calculation: If ReferenceAntigen is specified and ReferenceAntigenDilutionFactor is not Null or 1, automatically set to 1.1*ReferenceAntigenCoatingVolume/ReferenceAntigenDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
ReferenceAntigenDiluent
The buffer used to dilute the ReferenceAntigen as well as StandardReferenceAntigen and BlankReferenceAntigen to appropriate concentrations for the assay before plate coating. It provides conditions that support efficient antigen adsorption to the plate surface while preserving antigen stability.
Default Calculation: If reference antigens and their dilutionFactors for samples, standards and blanks are specified, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
ELISA Reagent Preparation
PrimaryAntibody
The antibody that directly binds to the TargetAntigen. It serves as the first layer of specific recognition and may be unlabeled or directly conjugated to a detection molecule (e.g., enzyme or fluorophore).
Default Calculation: For DirectELISA, DirectSandwichELISA, DirectCompetitiveELISA and FastELISA methods, automatically set to a labeled antibody against the TargetAntigen. For IndirectELISA, IndirectSandwichELISA and IndirectCompetitiveELISA, automatically set to an unlabeled antibody against the TargetAntigen. If no antibody is found against the TargetAntigen for direct methods, set to Model[Sample, "HRP-Conjugated Anti-Human-IL6 Antibody"]. If no antibody is found against the TargetAntigen for indirect methods, set to Model[Sample, "Monoclonal ANTI-FLAG M2 Antibody"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample.
Programmatic Pattern: (ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic
Index Matches to: experiment samples
PrimaryAntibodyDilutionFactor
The factor by which the PrimaryAntibody concentration is reduced by PrimaryAntibodyDiluent, calculated as the total final volume consisting of both the PrimaryAntibody and PrimaryAntibodyDiluent divided by the volume of the PrimaryAntibody. When PrimaryAntibodyDilutionFactor is 1, no dilution is performed. When preparing the primary antibody master mix in PrimaryAntibodyDilutionContainers, the total volume or prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same PrimaryAntibody and PrimaryAntibodyDilutionFactor. For example, if PrimaryAntibodyDilutionFactor is 1.25 and PrimaryAntibodyVolume is 100 Microliters, the primary antibody master mix is prepared by combining 100 Microliters of undiluted PrimaryAntibody with 25 Microliters of PrimaryAntibodyDiluent, assuming no duplicate dilutions are made. If three samples share the same PrimaryAntibody and its dilution factor, the master mix is prepared by mixing 300 Microliters of undiluted PrimaryAntibody with 75 Microliters of PrimaryAntibodyDiluent. This option is only applicable for DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA methods. For DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA methods, PrimaryAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA, automatically set to 1000.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: experiment samples
PrimaryAntibodyVolume
The volume of undiluted PrimaryAntibody added to PrimaryAntibodyDilutionContainers to prepare the diluted PrimaryAntibody by mixing with PrimaryAntibodyDiluent when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA, or the volume of undiluted PrimaryAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to mix with samples (both neat sample, spiked sample, and all diluted samples if applicable) at SampleVolume to form Sample-Antibody complex when Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA and PrimaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*PrimaryAntibodyImmunosorbentVolume/PrimaryAntibodyDilutionFactor. If Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA, automatically set to 2 Microliters.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
PrimaryAntibodyDilutionOnDeck
Indicates whether the PrimaryAntibody and PrimaryAntibodyDiluent are mixed on deck, immediately prior to being added to the ELISAPlate(s) during the immunosorbent step, instead of before the blocking step for DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA methods.
Default Calculation: Automatically set to False for DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA method when any PrimaryAntibodyDilutionFactor is specified.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
PrimaryAntibodyDiluent
The buffer used to dilute the PrimaryAntibody as well as StandardPrimaryAntibody and BlankPrimaryAntibody to their working concentrations. The diluent helps maintain the stability and activity of the antibody while minimizing nonspecific binding. This option is only applicable for DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA methods.
Default Calculation: If primary antibodies and their dilutionFactors for samples, standards and blanks are specified, automatically set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
IndirectELISA Reagent Preparation
SecondaryAntibody
The antibody that binds to the PrimaryAntibody rather than directly to the TargetAntigen. It is typically conjugated to a detection molecule (e.g., enzyme, fluorophore) to generate a measurable signal. This option is only applicable when Method is Indirect ELISA methods (IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA).
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA, automatically set to a stocked antibody against the PrimaryAntibody. If no antibody is found against the PrimaryAntibody for indirect methods, set to Model[Sample, "HRP-Conjugated Goat-Anti-Mouse-IgG Secondary Antibody"]
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: experiment samples
SecondaryAntibodyDilutionFactor
The factor by which the SecondaryAntibody concentration is reduced by SecondaryAntibodyDiluent, calculated as the total final volume consisting of both the SecondaryAntibody and SecondaryAntibodyDiluent divided by the volume of the SecondaryAntibody. When preparing the secondary antibody master mix in SecondaryAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same SecondaryAntibody and SecondaryAntibodyDilutionFactor. For example, if SecondaryAntibodyDilutionFactor is 1.25 and SecondaryAntibodyVolume is 100 Microliters, the secondary antibody master mix is prepared by combining 100 Microliters of undiluted SecondaryAntibody with 25 Microliters of SecondaryAntibodyDiluent, assuming no duplicate dilutions are made. If three samples share the same SecondaryAntibody and its dilution factor, the master mix is prepared by mixing 300 Microliters of undiluted SecondaryAntibody with 75 Microliters of SecondaryAntibodyDiluent.
Default Calculation: If SecondaryAntibody is specified, automatically set to 1000.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: experiment samples
SecondaryAntibodyVolume
The volume of undiluted SecondaryAntibody added to SecondaryAntibodyDilutionContainers to prepare the diluted SecondaryAntibody by mixing with SecondaryAntibodyDiluent corresponding to the input sample.
Default Calculation: If SecondaryAntibody is specified and SecondaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*SecondaryAntibodyImmunosorbentVolume/SecondaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: experiment samples
SecondaryAntibodyDilutionOnDeck
Indicates whether the SecondaryAntibody and SecondaryAntibodyDiluent are mixed on deck, immediately prior to being added to the ELISAPlate(s) during the immunosorbent step, instead of before the blocking step.
Default Calculation: Automatically set to False for indirect ELISA methods when any SecondaryAntibodyDilutionFactor is specified.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SecondaryAntibodyDiluent
The buffer used to dilute SecondaryAntibody as well as StandardSecondaryAntibody and BlankSecondaryAntibody to their working concentrations. The diluent helps maintain the stability and activity of the antibody while minimizing nonspecific binding.
Default Calculation: If secondary antibodies and their dilutionFactors for samples, standards and blanks are specified, automatically set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Plate Washing
WashingBuffer
The solution used to rinse off unbound coating reagents, samples and antibodies from the ELISAPlate(s). Washing is automatically performed by the plate washer integrated into the robotic Instrument after each assay step. The washing operation can be disabled for specific steps using the options CoatingWashing, BlockingWashing, PrimaryAntibodyImmunosorbentWashing, SecondaryAntibodyImmunosorbentWashing, SampleAntibodyComplexImmunosorbentWashing, and SampleImmunosorbentWashing.
Default Calculation: If any washing operation is required (at least one of the options CoatingWashing, BlockingWashing, PrimaryAntibodyImmunosorbentWashing, SecondaryAntibodyImmunosorbentWashing, SampleAntibodyComplexImmunosorbentWashing, SampleImmunosorbentWashing is True), automatically set to Model[Sample, StockSolution, "Phosphate Buffered Saline with 0.05% TWEEN 20, pH 7.4"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
WashPlateMethod
The configuration file that defines how the plate washer aspirates, dispenses, and manages liquid flow during ELISA washing steps, including speeds, heights, delays, and positioning for optimal washing efficiency. If your workflow requires parameters that are not currently supported by the predefined WashPlateMethod files, you can create and validate a custom method file using the WashPlate unit operation within ExperimentRoboticSamplePreparation beforehand. This option is only applicable when using the Instrument Model[Instrument, LiquidHandler, "Super STAR"] integrated with Model[Instrument, PlateWasher, "BioTek 405LS Microplate Washer"].
Default Calculation: Automatically set to Object[Method, WashPlate, "BioTek 405LS Default"] if Instrument is set to Model[Instrument, LiquidHandler, "Super STAR"] and WashingBuffer is not Null.
Pattern Description: An object of type or subtype Object[Method, WashPlate] or Null.
Programmatic Pattern: (ObjectP[{Object[Method, WashPlate]}] | Automatic) | Null
Blocking
BlockingBuffer
The solution used to prevent non-specific binding of antigen or antibody to the surface of the ELISAPlate(s). Blocking buffers typically contain inert proteins (e.g., BSA, casein, FBS) or specialized formulations that minimize background signal and enhance assay specificity.
Default Calculation: If Blocking is True, automatically set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlockingVolume
The amount of BlockingBuffer that is dispensed into the corresponding wells of the ELISAPlate(s), in order to prevent non-specific binding.
Default Calculation: If Blocking is True, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: experiment samples
BlockingTime
The duration of time when the BlockingBuffer is kept with the ELISAPlate(s), in order to prevent non-specific binding.
Default Calculation: If Blocking is True, automatically set to 1 Hour.
Pattern Description: Greater than or equal to 0 hours and less than or equal to 20 hours or Null.
Programmatic Pattern: (RangeP[0*Hour, 20*Hour] | Automatic) | Null
BlockingTemperature
The temperature at which the ELISAPlate(s) are kept during blocking, in order for the blocking analytes to be adsorbed to the unoccupied sites.
Default Calculation: If Blocking is True, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
BlockingMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during blocking. Gentle orbital shaking ensures even distribution of the blocking reagents. Shaking too vigorously, however, can cause splashing or evaporation, leading to inconsistent results.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
BlockingWashing
Indicates if a final washing step is performed at the end of BlockingTime before the immunosorbent steps. Block washing is generally recommended. Some specialized commercial pre-blocked plates may integrate blocking and sample addition without an intervening wash.
Default Calculation: If Blocking is set to True, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
BlockingWashVolume
The volume of WashBuffer added per wash cycle per well to rinse off the unbound blocking reagents from the ELISAPlate(s).
Default Calculation: If BlockingWashing is True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlockingNumberOfWashes
The number of washes performed to rinse off unbound blocking reagents. Each wash cycle first aspirates from, and then dispenses WashBuffer to, the ELISAPlate(s).
Default Calculation: If BlockingWashing is True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
PreImmunosorbent Step
PreImmunosorbentLigand
The reagent, such as a small molecule, receptor protein, or biologic drug, that binds to the TargetAntigen in input samples, standards, and blanks, enabling competitive or neutralization-based potency measurements. Unlike Spike or detection antibody, which serve as TargetAntigen or signal-generating detection label respectively, the PreImmunosorbentLigand is a defined binding partner added in a fixed, known quantity, to structurally alter the TargetAntigen. For DirectELISA and IndirectELISA method, the ligand is to be added to the ELISAPlates directly after blocking prior to immunosorbent steps to bind to the immunosorbent surface. For all other methods, the ligand is to be pre-mixed with the input samples, standards, and blanks on SampleAssemblyPlates prior to their transfer to ELISAPlates.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: (ObjectP[{Model[Sample], Object[Sample]}] | _String) | Null
PreImmunosorbentLigandVolume
The amount of PreImmunosorbentLigand added to either the ELISAPlates after blocking (for DirectELISA and IndirectELISA methods) or to the SampleAssemblyPlates during pre-mixing with input samples, standards, and blanks (for all other methods), to achieve the optimal binding interaction with the TargetAntigen for competitive or neutralization-based potency measurements.
Default Calculation: If PreImmunosorbentLigand is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
PreImmunosorbentIncubationTime
The duration for which the PreImmunosorbentLigand is incubated to allow sufficient binding interaction with the TargetAntigen, either on the ELISAPlates after blocking (for DirectELISA and IndirectELISA methods) or on the SampleAssemblyPlates during pre-mixing with input samples, standards, and blanks (for all other methods), prior to the immunosorbent steps.
Default Calculation: If PreImmunosorbentLigand is specified, automatically set to 15 minutes.
Pattern Description: Greater than or equal to 0 hours and less than or equal to 20 hours or Null.
Programmatic Pattern: (RangeP[0*Hour, 20*Hour] | Automatic) | Null
PreImmunosorbentIncubationTemperature
The temperature at which the PreImmunosorbentLigand is incubated during the PreImmunosorbentIncubationTime to ensure optimal binding interaction with the TargetAntigen, either on the ELISAPlates after blocking (for DirectELISA and IndirectELISA methods) or on the SampleAssemblyPlates during pre-mixing with input samples, standards, and blanks (for all other methods), prior to the immunosorbent steps.
Default Calculation: If PreImmunosorbentLigand is specified, automatically set to Ambient.
Pattern Description: Ambient or Custom or Null.
Programmatic Pattern: ((Ambient | RangeP[4*Celsius, 50*Celsius]) | Automatic) | Null
PreImmunosorbentIncubationMixRate
The speed at which the ELISAPlates (for DirectELISA and IndirectELISA methods) or SampleAssemblyPlates (for all other methods) are shaken (orbitally at a radius of 2 mm) during the PreImmunosorbentIncubationTime to ensure uniform and optimal binding interaction. Gentle orbital shaking ensures even distribution of the PreImmunosorbentLigand and consistent binding interaction with the TargetAntigen. Shaking too vigorously, however, can cause splashing or evaporation, leading to inconsistent results.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
PreImmunosorbentWashing
Indicates if a final washing step is performed at the end of the PreImmunosorbent incubation to remove unbound PreImmunosorbentLigand from the ELISAPlate(s) before the immunosorbent steps. This option is only applicable when Method is DirectELISA or IndirectELISA and PreImmunosorbentLigand is specified.
Default Calculation: If PreImmunosorbentLigand is specified and Method is DirectELISA or IndirectELISA, automatically set to True.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
PreImmunosorbentWashVolume
The volume of WashingBuffer added per wash cycle per well to rinse off unbound PreImmunosorbentLigand from the ELISAPlate(s).
Default Calculation: If PreImmunosorbentWashing is True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
PreImmunosorbentNumberOfWashes
The number of washes performed to rinse off unbound PreImmunosorbentLigand. Each wash cycle first aspirates from, and then dispenses WashingBuffer to, the ELISAPlate(s).
Default Calculation: If PreImmunosorbentWashing is True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
Detection
SubstrateSolution
The solution containing a chromogenic, fluorogenic, or chemiluminescent reagent that reacts with the enzyme conjugated to the detection antibody (or other enzyme label) to produce a measurable signal, enabling quantification of the target analyte in the assay. SubstrateSolution is sometimes referred to SubstrateSolutionA in a two-part substrate system, where SubstrateSolutionB contains an enhancer or activator that optimizes or initiates the reaction in SubstrateSolutionA.
Default Value: Model[Sample, ELISA TMB Stabilized Chromogen]
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample.
Programmatic Pattern: ObjectP[{Model[Sample], Object[Sample]}] | _String
SecondarySubstrateSolution
The solution containing an enhancer or activator that optimizes or initiates the reaction of SubstrateSolution when the two are mixed together. SecondarySubstrateSolution is sometimes referred to SubstrateSolutionB in a two-part substrate system.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: (ObjectP[{Model[Sample], Object[Sample]}] | _String) | Null
PreMixSubstrateSolution
Indicates whether the SubstrateSolution and SecondarySubstrateSolution are mixed before the blocking step instead of prior to being added to the ELISAPlate(s). If set to False, the substrate solutions are kept separate and combined into a master mix just before dispensing to the ELISAPlate(s). PreMixSubstrateSolution option is only applicable for two-part substrate systems.
Default Calculation: Automatically set to False if a SecondarySubstrateSolution is specified.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SubstrateSolutionMixRatio
The volume ratio of SubstrateSolution to SecondarySubstrateSolution. For example, when this option is set to 1, equal volumes of SubstrateSolution (SubstrateSolutionA) and SecondarySubstrateSolution (SubstrateSolutionB) are mixed to create the working substrate solution.
Default Calculation: Automatically set to 1 if a SecondarySubstrateSolution is specified.
Pattern Description: Greater than 0 or Null.
Programmatic Pattern: (GreaterP[0] | Automatic) | Null
SubstrateSolutionVolume
The volume of the working SubstrateSolution (pre-mixed according to the SubstrateSolutionMixRatio if a SecondarySubstrateSolution is specified) to be added to the assay plate(s). For example, when SubstrateSolutionVolume is set to 100 Microliters and SubstrateSolutionMixRatio is set to 1, 50 Microliters original SubstrateSolution and 50 Microliters SecondarySubstrateSolution is pre-mixed and added to each well on assay plate(s).
Default Value: 100 microliters
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters.
Programmatic Pattern: RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume]
SubstrateIncubationTime
The time allowed for the enzyme substrate reaction to occur in the ELISAPlate(s) before the reaction is stopped or measured.
Default Value: 30 minutes
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours.
Programmatic Pattern: RangeP[0*Minute, 24*Hour]
SubstrateIncubationTemperature
The temperature at which the ELISAPlate(s) are kept during SubstrateIncubation, in order for the detection reagent to react with antibody-conjugated enzyme.
Pattern Description: Ambient or Custom.
Programmatic Pattern: Ambient | RangeP[4*Celsius, 50*Celsius]
SubstrateIncubationMixRate
The speed at which the ELISAPlate(s) are shaken (orbitally, at a radius of 2 mm) during SubstrateIncubation.
Pattern Description: Greater than or equal to 40 revolutions per minute and less than or equal to 2500 revolutions per minute or Null.
Programmatic Pattern: RangeP[40*RPM, 2500*RPM] | Null
DetectionWashing
Indicates if an initial washing step is added to the detection step before adding SubstrateSolution. The DetectionWashingBuffer can be different from WashingBuffer used in Coating, Blocking and Immunosorbent steps. To prevent buffer interference with the following enzyme substrate reaction, the DetectionWashingBuffer usually does not contain detergents.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
DetectionWashingBuffer
The solution used to wash ELISAPlate(s) prior to adding SubstrateSolution. To prevent buffer interference with the following enzyme substrate reaction, the DetectionWashingBuffer usually does not contain detergents.
Default Calculation: If DetectionWashing is True, automatically set to Model[Sample, StockSolution, "1x PBS from 10X stock"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
DetectionWashVolume
The volume of DetectionWashingBuffer added per wash cycle per well to the assay plate(s).
Default Calculation: If DetectionWashing is set to True, automatically set to 250 Microliters.
Pattern Description: Greater than or equal to 50 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[Experiment`Private`$MinWashVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
DetectionNumberOfWashes
The number of washes performed for DetectionWashing. Each wash cycle first aspirates from, and then dispenses DetectionWashingBuffer to, the ELISAPlate(s).
Default Calculation: If DetectionWashing is set to True, automatically set to 4.
Pattern Description: Greater than or equal to 1 and less than or equal to 10 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 10, 1] | Automatic) | Null
RetainCover
Indicates if the lid(s) on the ELISAPlate(s) should not be taken off during measurement to decrease evaporation.
Pattern Description: True or False.
Programmatic Pattern: BooleanP
ReadDirection
Indicates the path the instrument follows when measuring signal (absorbance, fluorescence, or luminescence) across the wells of a plate. For example, when set to Column, the instrument reads all wells within a column before proceeding to the next column. Column, Row, SerpentineRow, SerpentineColumn are avaialbe for the "CLARIOstar" integrated to Model[Instrument, LiquidHandler, "Super STAR"], while only Column is available for "Reader96" integrated to Model[Instrument, LiquidHandler, "Hamilton Nimbus HD"].
Pattern Description: Row, Column, SerpentineRow, or SerpentineColumn.
Programmatic Pattern: ReadDirectionP
AbsorbanceDetection
StopSolution
The reagent that is used to stop colorimetric reaction between the enzyme and its substrate. This option is only applicable when DetectionMode is AbsorbanceIntensity.
Default Calculation: If DetectionMode is AbsorbanceIntensity and SubstrateSolution is Model[Sample, "ELISA TMB Stabilized Chromogen"], automatically set to Model[Sample, "ELISA HRP-TMB Stop Solution"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
StopSolutionVolume
The volume of StopSolution to be added to the ELISAPlate(s).
Default Calculation: If DetectionMode is AbsorbanceIntensity and StopSolution is specified, automatically set to 100 Microliters.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
AbsorbanceWavelength
The wavelength used to detect the absorbance of light by the product of the detection reaction. This option is only applicable when DetectionMode is AbsorbanceIntensity.
Default Calculation: Automatically set to 450 Nanometer if DetectionMode is AbsorbanceIntensity.
Pattern Description: Multiple Fixed Wavelengths or Multiple Wavelengths or Single Fixed Wavelength or Single Wavelength or Null.
Programmatic Pattern: (((405*Nanometer | 450*Nanometer | 492*Nanometer | 620*Nanometer) | RangeP[220*Nanometer, 1000*Nanometer] | {(405*Nanometer | 450*Nanometer | 492*Nanometer | 620*Nanometer)..} | {RangeP[220*Nanometer, 1000*Nanometer]..}) | Automatic) | Null
PrereadBeforeStop
Indicate if colorimetric reactions between the enzyme and its substrate will be checked by the plate reader before the the StopSolution is added to terminate the reaction. This option is only applicable when DetectionMode is AbsorbanceIntensity and a StopSolution is specified.
Default Calculation: Automatically set to True if DetectionMode is AbsorbanceIntensity and either PrereadTimepoints or PrereadAbsorbanceWavelength is specified. Automatically set to False if DetectionMode is AbsorbanceIntensity and a StopSolution is specified.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
PrereadTimepoints
The list of time points when absorbance intensitied at each PreparedAbsorbanceWavelength during the Preread process (before the colorimetric reaction was terminated).
Default Calculation: If PrereadBeforeStop is True, automatically set to the half of the specified SubstrateIncubationTime.
Pattern Description: Greater than or equal to 0 minutes and less than or equal to 24 hours or list of one or more greater than or equal to 0 minutes and less than or equal to 24 hours entries or Null.
Programmatic Pattern: ((RangeP[0*Minute, 24*Hour] | {RangeP[0*Minute, 24*Hour]..}) | Automatic) | Null
PrereadAbsorbanceWavelength
The wavelength used to detect the absorbance of light during the Preread process (before the colorimetric reaction was terminated).
Default Calculation: If PrereadBeforeStop is True, automatically set to the same value as the option AbsorbanceWavelength.
Pattern Description: Multiple Fixed Wavelengths or Multiple Wavelengths or Single Fixed Wavelength or Single Wavelength or Null.
Programmatic Pattern: (((405*Nanometer | 450*Nanometer | 492*Nanometer | 620*Nanometer) | RangeP[220*Nanometer, 1000*Nanometer] | {(405*Nanometer | 450*Nanometer | 492*Nanometer | 620*Nanometer)..} | {RangeP[220*Nanometer, 1000*Nanometer]..}) | Automatic) | Null
SignalCorrection
Indicates if an absorbance reading that is used to eliminate the interference of background absorbance (such as from ELISAPlate material and dust) is used. The correction is done by subtracting the reading at SignalCorrectionWavelength from that at the AbsorbanceWavelength. This option is only applicable when DetectionMode is AbsorbanceIntensity.
Default Calculation: Automatically set to True if SignalCorrectionWavelength is specified and DetectionMode is AbsorbanceIntensity. Otherwise, automatically set to False if DetectionMode is AbsorbanceIntensity.
Pattern Description: True or False or Null.
Programmatic Pattern: (BooleanP | Automatic) | Null
SignalCorrectionWavelength
The wavelength for absorbance reading that is used to eliminate the interference of background absorbance (such as from ELISAPlate material and dust). This option is only applicable when DetectionMode is AbsorbanceIntensity.
Default Calculation: Automatically set to 620 Nanometer if SignalCorrection is True and DetectionMode is AbsorbanceIntensity.
Pattern Description: Single Fixed Wavelength or Single Wavelength or Null.
Programmatic Pattern: (((405*Nanometer | 450*Nanometer | 492*Nanometer | 620*Nanometer) | RangeP[220*Nanometer, 1000*Nanometer]) | Automatic) | Null
AlternativeDetections
WavelengthSelection
Indicates if the emission wavelengths should be obtained by filters or monochromators.
Default Calculation: When EmissionWavelength(s) are set to specific wavelengths, automatically set to Monochromators provided all of the requested wavelengths, otherwise set to Filters. If EmissionWavelength is set to NoFilter when DetectionMode is LuminescenceIntensity, automatically set to NoFilter for WavelengthSelection as well.
Pattern Description: NoFilter, Filters, or Monochromators or Null.
Programmatic Pattern: (LuminescenceWavelengthSelectionP | Automatic) | Null
ReadLocation
Indicates if fluorescence or luminescence should be measured using an optic above the plate or one below the plate.
Default Calculation: Defaults to Bottom if RetainCover is set to True, otherwise defaults to Top.
Pattern Description: Top or Bottom or Null.
Programmatic Pattern: (ReadLocationP | Automatic) | Null
EmissionWavelength
The wavelength(s) at which fluorescence or luminescence emitted from the sample should be measured.
Default Calculation: When DetectionMode is LuminescenceIntensity, automatically set to to NoFilter. And when DetectionMode is FluorescenceIntensity, automatically set to 420 Nanometers.
Pattern Description: Greater than or equal to 320 nanometers and less than or equal to 740 nanometers or NoFilter or Null.
Programmatic Pattern: ((Null | (RangeP[320*Nanometer, 740*Nanometer] | NoFilter)) | Automatic) | Null
Index Matches to: EmissionWavelength
Gain
The gain which should be applied to the signal reaching the primary detector. This may be specified either as a direct voltage, or as a percentage (which indicates that the gain should be set such that the AdjustmentSample intensity is at the specified percentage of the instrument's dynamic range).
Default Calculation: Automatically set to 90% when DetectionMode is not AbsorbanceIntensity.
Pattern Description: Greater than or equal to 1 microvolt and less than or equal to 4095 microvolts or greater than or equal to 1 percent and less than or equal to 95 percent or Null.
Programmatic Pattern: ((RangeP[1*Percent, 95*Percent] | RangeP[1*Microvolt, 4095*Microvolt]) | Automatic) | Null
Index Matches to: EmissionWavelength
AdjustmentSample
The sample which should be used to perform automatic adjustments of gain and/or focal height values at run time. The focal height will be set so that the highest signal-to-noise ratio can be achieved for the AdjustmentSample. The gain will be set such that the AdjustmentSample fluoresces at the specified percentage of the instrument's dynamic range. When multiple aliquots of the same sample is used in the experiment, an index can be specified to use the desired aliquot for adjustments. When set to FullPlate, all wells of the assay plate are scanned and the well of the highest fluorescence intensity if perform gain and focal height adjustments.
Default Calculation: When DetectionMode is not AbsorbanceIntensity, automatically set to Individual if FocalHeight is set to Auto where the standard with the highest concentration is used to determine the gain percentage and focal height adjustment, otherwise set to FullPlate where all wells of the assay plate are scanned and the well of the highest fluorescence or luminescence intensity is used to determine the gain percentage and focal height adjustment.
Pattern Description: FullPlate or Individual or Null.
Programmatic Pattern: ((FullPlate | Individual) | Automatic) | Null
Index Matches to: EmissionWavelength
FocalHeight
The distance from the bottom of the plate carrier to the focal point where light from the sample is directed onto the detector. If set to Auto the height which corresponds to the highest AdjustmentSample signal will be used.
Default Calculation: If an adjustment sample is provided the height which corresponds to the highest AdjustmentSample signal will be used (as indicated by Auto). Otherwise defaults to 7 Millimeter.
Pattern Description: Auto or greater than or equal to 0 millimeters and less than or equal to 25 millimeters or Null.
Programmatic Pattern: ((Null | (RangeP[0*Millimeter, 25*Millimeter] | Auto)) | Automatic) | Null
Index Matches to: EmissionWavelength
FluorescenceDetection
ExcitationWavelength
For required EmissionWavelength, the wavelength of light used to excite the samples.
Default Calculation: When DetectionMode is FluorescenceIntensity, automatically set to 320 Nanometers.
Pattern Description: Greater than or equal to 320 nanometers and less than or equal to 740 nanometers or Null.
Programmatic Pattern: (RangeP[320*Nanometer, 740*Nanometer] | Automatic) | Null
Index Matches to: EmissionWavelength
LuminescenceDetection
IntegrationTime
The amount of time over which luminescence measurements should be integrated. Select a higher time to increase the reading intensity.
Default Calculation: Automatically set to 1 Second when DetectionMode is LuminescenceIntensity.
Pattern Description: Greater than or equal to 0.01 seconds and less than or equal to 100 seconds or Null.
Programmatic Pattern: ((Null | RangeP[0.01*Second, 100*Second]) | Automatic) | Null
Standard
Standard
A sample containing known amount of TargetAntigen molecule, used to generate a calibration curve for quantifying the TargetAntigen in experimental samples.
Default Calculation: If TargetAntigen is specified, automatically set SampleModel to the DefaultSampleModel of the TargetAntigen whenever either a non-precoated plate is used with DirectELISA or IndirectELISA, or the Method is any ELISA type other than DirectELISA or IndirectELISA.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardTargetAntigen
The analyte molecule(e.g., peptide, protein, or hormone) detected and quantified in the Standard samples by antibodies in the ELISA experiment. It should be the same as the TargetAntigen for all samples or a subset of samples to make the standard valid.
Default Calculation: Automatically set to the first Model[Molecule, Protein] in the Analytes field of the standard, or, if not populated, to the first Model[Molecule, Protein] in the Composition field of the standard, or if none exist, set to Null.
Pattern Description: An object of type or subtype Model[Molecule] or Null.
Programmatic Pattern: (ObjectP[{Model[Molecule]}] | Automatic) | Null
Index Matches to: Standard
StandardDilutionType
Indicates whether the standard is diluted, and if so, specifies the dilution type (linear or serial). Linear dilution represents a single stage dilution of the StandardTargetAntigen in the standard by specified StandardDilutionTransferVolume and StandardDilutionDiluentVolume. For example, a two-fold, five-fold and ten-fold linear dilution takes 500 Microliters, 200 Microliters and 100 Microliters of the original standard sample and generates three final diluted standards at 1 Milliliter (set by StandardTotalDilutionVolume at 1 Milliliter, StandardCumulativeDilutionFactor at {2, 5, 10} and StandardNumberOfDilutions at 3). Serial dilution represents a stepwise dilution of StandardTargetAntigen in the standard resulting in multiple standards and a geometric progression of the concentration. The first source standard is the original standard sample provided. For example, a standard sample with initial StandardTargetAntigen concentration of 100ng/ul, if diluted 6 times (set by StandardNumberOfDilutions at 6) and two-fold dilution at each stage (set by StandardSerialDilutionFactor at 2), should yield ELISA measurements of 50ng/ul, 25ng/ul, 12.5ng/ul, 6.25ng/ul, 3.125ug/ul, and 1.56ng/ul when StandardDilutionStrategy is Series or just 1.56ng/ul when StandardDilutionStrategy is Endpoint. This option is only applicable when a non-precoated plate is used with DirectELISA or IndirectELISA, or the Method is any ELISA type other than DirectELISA or IndirectELISA.
Default Calculation: Automatically set to Serial if Standard is specified and whenever either a non-precoated plate is used with DirectELISA or IndirectELISA, or the Method is any ELISA type other than DirectELISA or IndirectELISA.
Pattern Description: {Serial, Linear} or None or Null.
Programmatic Pattern: ((DilutionTypeP | None) | Automatic) | Null
Index Matches to: Standard
StandardDilutionStrategy
Indicates if only the final diluted standard (Endpoint) or all diluted standards (Series) produced by serial dilution are used in ELISA. This option is only applicable when StandardDilutionType is Serial.
Default Calculation: Automatically set to Series if StandardDilutionType is Serial.
Pattern Description: Series or Endpoint or Null.
Programmatic Pattern: ((Null | DilutionStrategyP) | Automatic) | Null
Index Matches to: Standard
StandardNumberOfDilutions
The number of diluted standards to prepare. When StandardDilutionStrategy is Linear, all prepared diluted standards are directly diluted from the undiluted Standard but can be different in StandardCumulativeDilutionFactor. When StandardDilutionStrategy is Serial, the dilution is performed stepwise, producing standards with progressively decreasing concentrations across all dilution levels.
Default Calculation: Automatically set to 1 when StandardDilutionStrategy is Linear, and automatically set to the length of StandardCumulativeDilutionFactor, or StandardSerialDilutionFactor if provided when StandardDilutionStrategy is Serial, or 6 if not provided.
Pattern Description: Greater than or equal to 1 and less than or equal to 15 in increments of 1 or Null.
Programmatic Pattern: (RangeP[1, 15, 1] | Automatic) | Null
Index Matches to: Standard
StandardCumulativeDilutionFactor
The factor by which the concentration of the StandardTargetAntigen in the original standard is reduced during the dilution. When StandardDilutionType is Serial, the length of this list must match the corresponding value in StandardNumberOfDilutions. In the example in the option StandardDilutionType when it is set to Serial, the two-fold dilution for 6 times is corresponding to StandardCumulativeDilutionFactor at {2,4,8,16,32,64}. When StandardDilutionType is Linear, each dilution is made independently from the original standard. In the example in the option StandardDilutionType when it is set to Linear, a two-fold, five-fold and ten-fold linear dilution is corresponding to StandardCumulativeDilutionFactor at {2,5,10}.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 1 and less than or equal to 100000000000000000000000 or Null.
Programmatic Pattern: (RangeP[1, 10^23] | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardSerialDilutionFactor
The factor by which the concentration of the StandardTargetAntigen in the resulting standard of the previous dilution step is reduced. For example, if the StandardCumulativeDilutionFactor is equal to {10,100,1000}, StandardSerialDilutionFactor is 10 or {10,10,10}. Because the dilution curve does not intrinsically include the original (undiluted) standard, the first dilution factor must be set to 1 if the original standard is to be measured as well. Using the same example described in StandardDilutionType, to obtain ELISA measurements of 100 ng/ul, 50ng/ul, 25ng/ul, 12.5ng/ul, 6.25ng/ul, 3.125ug/ul, and 1.56ng/ul, StandardSerialDilutionFactor should be defined as {1, 2, 2, 2, 2, 2, 2}, and StandardNumberOfDilutions should be set to 7. This option is only applicable when StandardDilutionType is Serial.
Default Calculation: Automatically set based on the systems of equations specified under the StandardDilutionType option. If no standard dilution options are specified, automatically set StandardSerialDilutionFactor to 2 when StandardDilutionStrategy is Serial.
Pattern Description: Greater than or equal to 1 and less than or equal to 100000000000000000000000 or Null.
Programmatic Pattern: ((Null | RangeP[1, 10^23]) | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardDilutionTransferVolume
The amount of standard that is diluted with StandardDiluent when StandardDilutionType is set to Linear, or the amount of standard transferred from the resulting standard of one round of the dilution series to the next standard in the series when StandardDilutionType is set to Serial.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardDilutionDiluentVolume
The amount of StandardDiluent added to dilute the standard when StandardDilutionType is set to Linear, or the amount of StandardDiluent added to dilute the standard at each stage of the dilution when StandardDilutionType is set to Serial.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: ((Null | RangeP[0*Milliliter, $MaxTransferVolume]) | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardTotalDilutionVolume
The total volume of Standard and StandardDiluent. If StandardDilutionType is set to Serial, this is also the volume of the resulting standard before StandardDilutionTransferVolume has been removed for use in the next dilution in the series. If StandardDilutionType is set to Linear, this is the total amount of StandardDilutionDiluentVolume and StandardDilutionTransferVolume.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardDilutionFinalVolume
The volume of the resulting diluted standard after StandardDilutionTransferVolume has been removed for use in the next dilution in the series if StandardDilutionType is set to Serial. This option is only applicable when StandardDilutionType is Serial.
Default Calculation: Automatically set based on the systems of equations specified under the DilutionType option.
Pattern Description: Greater than or equal to 0 milliliters and less than or equal to 20 liters or Null.
Programmatic Pattern: (RangeP[0*Milliliter, $MaxTransferVolume] | Automatic) | Null
Index Matches to: Standard
Nested Index Matches to: Standard
StandardDestinationWell
The well locations assigned for each output standard, corresponding to the given input standard. This includes all diluted series if applicable. If more than one ELISAPlate is utilized, the standard is place on the same well position on each plate. When configuring StandardDestinationWell, set IndexMatch to Variable and ensure the number of DestinationWell entries exactly matches the total number of standard SamplesOut. Please use ExperimentELISAPreview to check the arrangement of Standards on the ELISAPlates.
Default Calculation: If Method is DirectELISA or IndirectELISA and Coating is False, automatically set to the well and plate position of the input standard. Otherwise, filling the ELISAPlate(s) in the order of MoatingBuffer and Blank first, then starting the well assignment for standards from the first available well, and continue columnwise.
Pattern Description: List of one or more A1, A2, A3, A4, A5, A6, A7, A8, A9, A10, A11, A12, B1, B2, B3, B4, B5, B6, B7, B8, B9, B10, B11, B12, C1, C2, C3, C4, C5, C6, C7, C8, C9, C10, C11, C12, D1, D2, D3, D4, D5, D6, D7, D8, D9, D10, D11, D12, E1, E2, E3, E4, E5, E6, E7, E8, E9, E10, E11, E12, F1, F2, F3, F4, F5, F6, F7, F8, F9, F10, F11, F12, G1, G2, G3, G4, G5, G6, G7, G8, G9, G10, G11, G12, H1, H2, H3, H4, H5, H6, H7, H8, H9, H10, H11, or H12 entries or Null.
Programmatic Pattern: ({Alternatives @@ Flatten[AllWells[NumberOfWells -> 96]]..} | Automatic) | Null
Index Matches to: Standard
StandardDiluent
The buffer used to perform multiple dilutions on Standards to appropriate concentrations.
Default Calculation: If StandardDilutionType is not None or Null for all standards, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"] when Method is set to DirectELISA or IndirectELISA, otherwise set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
StandardVolume
The volume of prepared standard(s) (all diluted standards if applicable) that are dispensed into the SampleAssemblyPlate(s) in order to form Standard-Antibody complex when Method is FastELISA, DirectCompetitiveELISA, or IndirectCompetitiveELISA.
Default Calculation: If Method is FastELISA, DirectCompetitiveELISA, or IndirectCompetitiveELISA and Standard is not Null, automatically set to the same value as the option SampleVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardCoatingVolume
The amount of prepared standard(s) that are dispensed into the ELISAPlate(s), in order for the Standard to be adsorbed to the surface of the well.
Default Calculation: If Standard is not Null and Method is DirectELISA or IndirectELISA, automatically set to the same value as the option SampleCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardImmunosorbentVolume
The volume of the prepared standard(s) to be loaded on the ELISAPlate for the StandardTargetAntigen to bind to the StandardCaptureAntibody in DirectSandwichELISA and IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA and Standard is specified, automatically set to the same value as the option SampleImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardCoatingAntibody
The antibody to be immobilized on the surface of the ELISA plate(s) to capture the StandardTargetAntigen during the assay. This option is only applicable when Method is FastELISA. It should be the same as the CoatingAntibody for all samples or a subset of samples to make the standard valid.
Default Calculation: If Method is FastELISA and Standard is not Null, automatically set to the same value in the option CoatingAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardCoatingAntibodyDilutionFactor
The factor by which the StandardCoatingAntibody concentration is reduced by CoatingAntibodyDiluent, calculated as the total final volume consisting of both the StandardCoatingAntibody and CoatingAntibodyDiluent divided by the volume of the StandardCoatingAntibody. When StandardCoatingAntibodyDilutionFactor is 1, no dilution is performed. When preparing the coating antibody master mix in CoatingAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same CoatingAntibody and CoatingAntibodyDilutionFactor. For example, if StandardCoatingAntibodyDilutionFactor is 100 and StandardCoatingAntibodyVolume is 5 Microliters, the coating antibody master mix is prepared by combining 5 Microliters of undiluted StandardCoatingAntibody with 495 Microliters of CoatingAntibodyDiluent. If the input sample shares the same CoatingAntibody and its dilution factor with the standard, the master mix is prepared by mixing 10 Microliters of undiluted CoatingAntibody with 990 Microliters of CoatingAntibodyDiluent. This option is only applicable when Method is FastELISA.
Default Calculation: If Method is FastELISA and Standard is specified, automatically set to the same value in the option CoatingAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: Standard
StandardCoatingAntibodyVolume
The volume of undiluted StandardCoatingAntibody directly added into the CoatingAntibodyDilutionContainers to mix with CoatingAntibodyDiluent. This option is only applicable when StandardCoatingAntibodyDilutionFactor is not Null or 1 and StandardCoatingAntibody is specified.
Default Calculation: If StandardCoatingAntibody is specified and StandardCoatingAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*StandardCoatingAntibodyCoatingVolume/StandardCoatingAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: Standard
StandardCoatingAntibodyCoatingVolume
The amount of StandardCoatingAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the StandardCoatingAntibody to be adsorbed to the surface of the well.
Default Calculation: If Method is FastELISA and Standard is not Null, automatically set to the same value in the option CoatingAntibodyCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardCaptureAntibody
The antibody that is used to pull down the StandardTargetAntigen from standard solution to the surface of the ELISAPlate(s) in DirectSandwichELISA, IndirectSandwichELISA, and FastELISA methods. It should be the same as the CaptureAntibody for all samples or a subset of samples to make the standard valid.
Default Calculation: If Method is FastELISA, DirectSandwichELISA or IndirectSandwichELISA and Standard is specified, automatically set to the same value in the option CaptureAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardCaptureAntibodyDilutionFactor
The factory by which the StandardCaptureAntibody concentration is reduced by CaptureAntibodyDiluent, calculated as the total final volume consisting of both the StandardCaptureAntibody and CaptureAntibodyDiluent divided by the volume of the StandardCaptureAntibody. This option is only applicable for DirectSandwichELISA and IndirectSandwichELISA methods. For FastELISA method, StandardCaptureAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and Standard is specified, automatically set to the same value in the option CaptureAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: Standard
StandardCaptureAntibodyVolume
The volume of undiluted StandardCaptureAntibody added into the CaptureAntibodyDilutionContainers to mix with CaptureAntibodyDiluent when Method is DirectSandwichELISA or IndirectSandwichELISA, or the volume of undiluted StandardCaptureAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to form Standard-Antibody complex when Method is FastELISA.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and StandardCaptureAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*StandardCaptureAntibodyCoatingVolume/StandardCaptureAntibodyDilutionFactor if Standard is not Null and StandardCaptureAntibody is specified. If Method is FastELISA and Standard is specified, automatically set to the same value in the option CaptureAntibodyVolume.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: Standard
StandardCaptureAntibodyCoatingVolume
The amount of StandardCaptureAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the StandardCaptureAntibody to be adsorbed to the surface of the well. This option is only applicable when Method is DirectSandwichELISA or IndirectSandwichELISA.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and Standard is specified, automatically set to the same value in the option CaptureAntibodyCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardReferenceAntigen
The antigen that competes with StandardTargetAntigen in the standard for the binding of the StandardPrimaryAntibody. It is also referred to as inhibitor antigen, and should be the same as the ReferenceAntigen for all samples or a subset of samples to make the standard valid. This option is only applicable when Method is DirectCompetitiveELISA or IndirectCompetitiveELISA.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Standard is not Null, automatically set to the same value as the option ReferenceAntigen.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardReferenceAntigenDilutionFactor
The factor by which the StandardReferenceAntigen concentration is reduced by ReferenceAntigenDiluent, calculated as the total final volume consisting of both the StandardReferenceAntigen and ReferenceAntigenDiluent divided by the volume of the StandardReferenceAntigen. When StandardReferenceAntigenDilutionFactor is 1, no dilution is performed. When preparing the reference antigen master mix in ReferenceAntigenDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same ReferenceAntigen and ReferenceAntigenDilutionFactor. For example, if StandardReferenceAntigenDilutionFactor is 100 and StandardReferenceAntigenVolume is 5 Microliters, the reference antigen master mix is prepared by combining 5 Microliters of undiluted ReferenceAntigen with 495 Microliters of ReferenceAntigenDiluent, assuming no duplicate dilutions are made. If one input sample shares the same ReferenceAntigen and its dilution factor with the standard, the master mix is prepared by mixing 10 Microliters of undiluted ReferenceAntigen with 990 Microliters of ReferenceAntigenDiluent. This option is only applicable for DirectCompetitiveELISA or IndirectCompetitiveELISA methods.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Standard is not Null, automatically set to the same value as ReferenceAntigenDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: Standard
StandardReferenceAntigenVolume
The volume of undiluted StandardReferenceAntigen added into the ReferenceAntigenDilutionContainers to mix with ReferenceAntigenDiluent. This option is only applicable when StandardReferenceAntigenDilutionFactor is not Null or 1 and StandardReferenceAntigen is specified.
Default Calculation: If StandardReferenceAntigen is specified and StandardReferenceAntigenDilutionFactor is not Null or 1, automatically set to 1.1*StandardReferenceAntigenCoatingVolume/StandardReferenceAntigenDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardReferenceAntigenCoatingVolume
The amount of StandardReferenceAntigen (either diluted or undiluted) that is dispensed into the ELISAPlate(s), in order for the StandardReferenceAntigen to be adsorbed to the surface of the well.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Standard is not Null, automatically set to the same value as ReferenceAntigenCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardPrimaryAntibody
The antibody that directly binds to the StandardTargetAntigen. It should be the same as the PrimaryAntibody for all samples or a subset of samples to make the standard valid.
Default Calculation: If Standard is not Null, automatically set to an antibody against the StandardTargetAntigen or the first PrimaryAntibody for the input samples.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardPrimaryAntibodyDilutionFactor
The factor by which the StandardPrimaryAntibody concentration is reduced by PrimaryAntibodyDiluent, calculated as the total final volume consisting of both the StandardPrimaryAntibody and PrimaryAntibodyDiluent divided by the volume of the StandardPrimaryAntibody. When StandardPrimaryAntibodyDilutionFactor is 1, no dilution is performed. When preparing the primary antibody master mix in PrimaryAntibodyDilutionContainers, the total volume or prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same PrimaryAntibody and PrimaryAntibodyDilutionFactor. For example, if StandardPrimaryAntibodyDilutionFactor is 1.25 and StandardPrimaryAntibodyVolume is 100 Microliters, the primary antibody master mix is prepared by combining 100 Microliters of undiluted PrimaryAntibody with 25 Microliters of PrimaryAntibodyDiluent, assuming no duplicate dilutions are made. If one input sample shares the same PrimaryAntibody and its dilution factor with the standard, the master mix is prepared by mixing 200 Microliters of undiluted PrimaryAntibody with 50 Microliters of PrimaryAntibodyDiluent. This option is only applicable for DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA methods. For DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA methods, PrimaryAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: When Standard is not Null and used for DirectELISA, IndirectELISA, DirectSandwichELISA, IndirectSandwichELISA, automatically set to the same value as option PrimaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: Standard
StandardPrimaryAntibodyVolume
The volume of undiluted StandardPrimaryAntibody added to PrimaryAntibodyDilutionContainers to prepare the diluted StandardPrimaryAntibody by mixing with PrimaryAntibodyDiluent when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA, or the volume of undiluted StandardPrimaryAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to mix with standards (all diluted standards if applicable) at StandardVolume to form Standard-Antibody complex when Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA and StandardPrimaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*StandardPrimaryAntibodyImmunosorbentVolume/StandardPrimaryAntibodyDilutionFactor. If Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA and Standard is specified, automatically set to the same value as the option PrimaryAntibodyVolume.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: Standard
StandardPrimaryAntibodyImmunosorbentVolume
The volume of the StandardPrimaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for the Immunosorbent step when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA and Standard is specified, automatically set to the same value as PrimaryAntibodyImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardSecondaryAntibody
The antibody that binds to the StandardPrimaryAntibody rather than directly to the StandardTargetAntigen. It is typically conjugated to a detection molecule (e.g., enzyme, fluorophore) to generate a measurable signal. This option is only applicable when Method is Indirect ELISA methods (IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA). It should be the same as the SecondaryAntibody for all samples or a subset of samples to make the standard valid.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Standard is specified, automatically set to a stocked secondary antibody if StandardPrimaryAntibody is specified or the first SecondaryAntibody for the input samples.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
Index Matches to: Standard
StandardSecondaryAntibodyDilutionFactor
The factor by which the StandardSecondaryAntibody concentration is reduced by SecondaryAntibodyDiluent, calculated as the total final volume consisting of both the StandardSecondaryAntibody and SecondaryAntibodyDiluent divided by the volume of the StandardSecondaryAntibody. When preparing the secondary antibody master mix in SecondaryAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same SecondaryAntibody and SecondaryAntibodyDilutionFactor. For example, if StandardSecondaryAntibodyDilutionFactor is 1.25 and StandardSecondaryAntibodyVolume is 100 Microliters, the secondary antibody master mix is prepared by combining 100 Microliters of undiluted SecondaryAntibody with 25 Microliters of SecondaryAntibodyDiluent, assuming no duplicate dilutions are made. If one input sample shares the same SecondaryAntibody and its dilution factor with this standard, the master mix is prepared by mixing 200 Microliters of undiluted SecondaryAntibody with 50 Microliters of SecondaryAntibodyDiluent.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Standard is specified, automatically set to the same value as the option SecondaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
Index Matches to: Standard
StandardSecondaryAntibodyVolume
The volume of undiluted StandardSecondaryAntibody added to SecondaryAntibodyDilutionContainers to prepare the diluted StandardSecondaryAntibody by mixing with SecondaryAntibodyDiluent per standard.
Default Calculation: If StandardSecondaryAntibody is specified and StandardSecondaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*StandardSecondaryAntibodyImmunosorbentVolume/StandardSecondaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
Index Matches to: Standard
StandardSecondaryAntibodyImmunosorbentVolume
The volume of the StandardSecondaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for the immunosorbent step.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Standard is specified, automatically set to the same value as the option SecondaryAntibodyImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardBlockingVolume
The amount of BlockingBuffer that is dispensed into the corresponding wells of the ELISAPlate(s), in order to prevent non-specific binding.
Default Calculation: If Blocking is True and Standard is specified, automatically set to the same value as the option BlockingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
StandardAntibodyComplexImmunosorbentVolume
The volume of the standard-antibody complex to be loaded on the ELISAPlate(s). In DirectCompetitiveELISA and IndirectCompetitiveELISA, this step enables the free StandardPrimaryAntibody to bind to the StandardReferenceAntigen coated on the ELISAPlate(s). In FastELISA, this step enables the PrimaryAntibody-TargetAntigen-CaptureAntibody complex to bind to the StandardCoatingAntibody on the ELISAPlate(s).
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA and Standard is specified, automatically set to the same value as the option SampleAntibodyComplexImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Index Matches to: Standard
Blank
Blank
The solution containing no TargetAntigen, used as a baseline or negative control for the ELISA.
Default Calculation: If Method is DirectELISA or IndirectELISA and Coating is True, automatically set to Model[Sample, StockSolution, "1x Carbonate-Bicarbonate Buffer pH10"]. For other methods, automatically set to Model[Sample, "ELISA Blocker Blocking Buffer"].
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankTargetAntigen
The analyte molecule(e.g., peptide, protein, or hormone) used in the blanks as baseline or negative control. It should be the same as the TargetAntigen for all samples or a subset of samples to make the blank valid.
Default Calculation: Automatically match TargetAntigen of the input samples if Blank is specified.
Pattern Description: An object of type or subtype Model[Molecule] or Null.
Programmatic Pattern: (ObjectP[{Model[Molecule]}] | Automatic) | Null
BlankDestinationWell
The assigned well location for the input blank. If more than one ELISAPlate is utilized, the blank is place on the same well on each plate. Please use ExperimentELISAPreview to check the arrangement of Blanks on the ELISAPlates.
Default Calculation: If Method is DirectELISA or IndirectELISA and Coating is False, automatically set to the well and plate position of the input blank. Otherwise, filling the ELISAPlate(s) in the order of MoatingBuffer first, then starting the well assignment for blanks from the first available well, and continue columnwise.
Pattern Description: A1, A2, A3, A4, A5, A6, A7, A8, A9, A10, A11, A12, B1, B2, B3, B4, B5, B6, B7, B8, B9, B10, B11, B12, C1, C2, C3, C4, C5, C6, C7, C8, C9, C10, C11, C12, D1, D2, D3, D4, D5, D6, D7, D8, D9, D10, D11, D12, E1, E2, E3, E4, E5, E6, E7, E8, E9, E10, E11, E12, F1, F2, F3, F4, F5, F6, F7, F8, F9, F10, F11, F12, G1, G2, G3, G4, G5, G6, G7, G8, G9, G10, G11, G12, H1, H2, H3, H4, H5, H6, H7, H8, H9, H10, H11, or H12 or Null.
Programmatic Pattern: (Alternatives @@ Flatten[AllWells[NumberOfWells -> 96]] | Automatic) | Null
BlankVolume
The volume of Blank that is dispensed into the SampleAssemblyPlate(s) to mix with antibodies as a control for the Sample-Antibody complex formation when Method is FastELISA, DirectCompetitiveELISA, or IndirectCompetitiveELISA.
Default Calculation: If Method is FastELISA, DirectCompetitiveELISA, or IndirectCompetitiveELISA and Blank is not Null, automatically set to the same value as the option SampleVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankCoatingVolume
The amount of Blank that is dispensed into the ELISAPlate(s) as a control for sample coating when Method is DirectELISA or IndirectELISA.
Default Calculation: If Blank is not Null and Method is DirectELISA or IndirectELISA, automatically set to the same value as the option SampleCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankImmunosorbentVolume
The volume of the Blank to be loaded on the ELISAPlate(s) as a control for the target antigen to bind to the capture antibody in DirectSandwichELISA and IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectSandwichELISA and IndirectSandwichELISA and Blank is specified, automatically set to the same value as the option SampleImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankCoatingAntibody
The coating antibody added to the Blank wells of the ELISAPlate(s), serving as a coating control to match surface binding conditions of the Sample wells. This option is only applicable when Method is FastELISA. It should be the same as the CoatingAntibody for all samples or a subset of samples to make the blank valid.
Default Calculation: If Method is FastELISA and Blank is not Null, automatically set to the same value in the option CoatingAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankCoatingAntibodyDilutionFactor
The factor by which the BlankCoatingAntibody concentration is reduced by CoatingAntibodyDiluent, calculated as the total final volume consisting of both the BlankCoatingAntibody and CoatingAntibodyDiluent divided by the volume of the BlankCoatingAntibody. When BlankCoatingAntibodyDilutionFactor is 1, no dilution is performed. When preparing the coating antibody master mix in CoatingAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same CoatingAntibody and CoatingAntibodyDilutionFactor. For example, if BlankdCoatingAntibodyDilutionFactor is 100 and BlankCoatingAntibodyVolume is 5 Microliters, the coating antibody master mix is prepared by combining 5 Microliters of undiluted StandardCoatingAntibody with 495 Microliters of CoatingAntibodyDiluent. If the input sample shares the same CoatingAntibody and its dilution factor with the blank, the master mix is prepared by mixing 10 Microliters of undiluted CoatingAntibody with 990 Microliters of CoatingAntibodyDiluent. This option is only applicable when Method is FastELISA.
Default Calculation: If Method is FastELISA and Blank is specified, automatically set to the same value in the option CoatingAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
BlankCoatingAntibodyVolume
The volume of undiluted BlankCoatingAntibody directly added into the CoatingAntibodyDilutionContainers to mix with CoatingAntibodyDiluent. This option is only applicable when BlankCoatingAntibodyDilutionFactor is not Null or 1 and BlankCoatingAntibody is specified.
Default Calculation: If BlankCoatingAntibody is specified and BlankCoatingAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*BlankCoatingAntibodyCoatingVolume/BlankCoatingAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
BlankCoatingAntibodyCoatingVolume
The amount of BlankCoatingAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s).
Default Calculation: If Method is FastELISA and Blank is specified, automatically set to the same value in the option CoatingAntibodyCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankCaptureAntibody
The antibody that is added to the Blank wells of the ELISAPlate(s) or SampleAssemblyPlate(s), serving as a control for binding the target antigen in DirectSandwichELISA, IndirectSandwichELISA, and FastELISA methods. It should be the same as the CaptureAntibody for all samples or a subset of samples to make the blank valid.
Default Calculation: If Method is FastELISA, DirectSandwichELISA or IndirectSandwichELISA and Blank is specified, automatically set to the same value in the option CaptureAntibody.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankCaptureAntibodyDilutionFactor
The factory by which the BlankCaptureAntibody concentration is reduced by CaptureAntibodyDiluent, calculated as the total final volume consisting of both the BlankCaptureAntibody and CaptureAntibodyDiluent divided by the volume of the BlankCaptureAntibody. This option is only applicable for DirectSandwichELISA and IndirectSandwichELISA methods. For FastELISA method, BlankCaptureAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and Blank is specified, automatically set to the same value in the option CaptureAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
BlankCaptureAntibodyVolume
The volume of undiluted BlankCaptureAntibody added into the CaptureAntibodyDilutionContainers to mix with CaptureAntibodyDiluent when Method is DirectSandwichELISA or IndirectSandwichELISA, or the volume of undiluted BlankCaptureAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to mix with Blank during SampleAntibodyComplexIncubation when Method is FastELISA.
Default Calculation: If Method is DirectSandwichELISA or IndirectSandwichELISA and BlankCaptureAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*BlankCaptureAntibodyCoatingVolume/BlankCaptureAntibodyDilutionFactor. If Method is FastELISA and and Blank is not Null, automatically set to the same value in the option CaptureAntibodyVolume.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
BlankCaptureAntibodyCoatingVolume
The amount of BlankCaptureAntibody (either diluted or undiluted) that is dispensed into the ELISAPlate(s) during the coating step. This option is only applicable when Method is DirectSandwichELISA or IndirectSandwichELISA.
Default Calculation: If Blank is specified and Method is DirectSandwichELISA or IndirectSandwichELISA, automatically set to the same value in the option CaptureAntibodyCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankReferenceAntigen
The antigen that competes with BlankTargetAntigen in the blank for the binding of the BlankPrimaryAntibody. It is also referred to as inhibitor antigen. This option is only applicable when Method is DirectCompetitiveELISA or IndirectCompetitiveELISA. It should be the same as the ReferenceAntigen for all samples or a subset of samples to make the blank valid.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Blank is not Null, automatically set to the same value as the option ReferenceAntigen.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankReferenceAntigenDilutionFactor
The factor by which the BlankReferenceAntigen concentration is reduced by ReferenceAntigenDiluent, calculated as the total final volume consisting of both the BlankReferenceAntigen and ReferenceAntigenDiluent divided by the volume of the BlankReferenceAntigen. When BlankReferenceAntigenDilutionFactor is 1, no dilution is performed. When preparing the reference antigen master mix in ReferenceAntigenDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same ReferenceAntigen and ReferenceAntigenDilutionFactor. For example, if BlankReferenceAntigenDilutionFactor is 100 and BlankReferenceAntigenVolume is 5 Microliters, the reference antigen master mix is prepared by combining 5 Microliters of undiluted ReferenceAntigen with 495 Microliters of ReferenceAntigenDiluent, assuming no duplicate dilutions are made. If one input sample shares the same ReferenceAntigen and its dilution factor with the blank, the master mix is prepared by mixing 10 Microliters of undiluted ReferenceAntigen with 990 Microliters of ReferenceAntigenDiluent. This option is only applicable for DirectCompetitiveELISA or IndirectCompetitiveELISA methods.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Blank is not Null, automatically set to the same value as ReferenceAntigenDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
BlankReferenceAntigenVolume
The volume of undiluted BlankReferenceAntigen directly added into the ReferenceAntigenDilutionContainers to mix with ReferenceAntigenDiluent. This option is only applicable when BlankReferenceAntigenDilutionFactor is not Null or 1 and BlankReferenceAntigen is specified.
Default Calculation: If BlankReferenceAntigen is specified and BlankReferenceAntigenDilutionFactor is not Null or 1, automatically set to 1.1*BlankReferenceAntigenCoatingVolume/BlankReferenceAntigenDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankReferenceAntigenCoatingVolume
The amount of diluted BlankReferenceAntigen that is dispensed into the ELISAPlate(s) during the coating step.
Default Calculation: If Method DirectCompetitiveELISA or IndirectCompetitiveELISA and Blank is not Null, automatically set to the same value as ReferenceAntigenCoatingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankPrimaryAntibody
The antibody that directly binds with the BlankTargetAntigen. It should be the same as the PrimaryAntibody for all samples or a subset of samples to make the blank valid.
Default Calculation: If Blank is not Null, automatically set to an antibody against the BlankTargetAntigen or the first PrimaryAntibody for the input samples.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankPrimaryAntibodyDilutionFactor
The factor by which the BlankPrimaryAntibody concentration is reduced by PrimaryAntibodyDiluent, calculated as the total final volume consisting of both the BlankPrimaryAntibody and PrimaryAntibodyDiluent divided by the volume of the BlankPrimaryAntibody. When BlankPrimaryAntibodyDilutionFactor is 1, no dilution is performed. When preparing the primary antibody master mix in PrimaryAntibodyDilutionContainers, the total volume or prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same PrimaryAntibody and PrimaryAntibodyDilutionFactor. For example, if BlankPrimaryAntibodyDilutionFactor is 1.25 and BlankPrimaryAntibodyVolume is 100 Microliters, the primary antibody master mix is prepared by combining 100 Microliters of undiluted PrimaryAntibody with 25 Microliters of PrimaryAntibodyDiluent, assuming no duplicate dilutions are made. If one input sample shares the same PrimaryAntibody and its dilution factor with the blank, the master mix is prepared by mixing 200 Microliters of undiluted PrimaryAntibody with 50 Microliters of PrimaryAntibodyDiluent. This option is only applicable for DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA methods. For DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA methods, PrimaryAntibody dilution during the experiment is not permitted and must be performed beforehand.
Default Calculation: When Blank is not Null and used for DirectELISA, IndirectELISA, DirectSandwichELISA, IndirectSandwichELISA, automatically set to the same value as option PrimaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
BlankPrimaryAntibodyVolume
The volume of undiluted BlankPrimaryAntibody added to PrimaryAntibodyDilutionContainers to prepare the diluted BlankPrimaryAntibody by mixing with PrimaryAntibodyDiluent when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA, or the volume of undiluted BlankPrimaryAntibody directly added into the corresponding well of the SampleAssemblyPlate(s) to mix with the blank at BlankVolume during SampleAntibodyIncubation when Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, and FastELISA.
Default Calculation: If Method is DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA and BlankPrimaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*BlankPrimaryAntibodyImmunosorbentVolume/BlankPrimaryAntibodyDilutionFactor. If Method is DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA and Blank is specified, automatically set to the same value as the option PrimaryAntibodyVolume.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
BlankPrimaryAntibodyImmunosorbentVolume
The volume of the BlankPrimaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for the Immunosorbent step when Method is DirectELISA, IndirectELISA, DirectSandwichELISA, and IndirectSandwichELISA.
Default Calculation: If the Method is set to DirectELISA, IndirectELISA, DirectSandwichELISA, or IndirectSandwichELISA and Blank is specified, automatically set to the same value as PrimaryAntibodyImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankSecondaryAntibody
The antibody that binds to the BlankPrimaryAntibody rather than directly to the BlankTargetAntigen. It is typically conjugated to a detection molecule (e.g., enzyme, fluorophore) to generate a measurable signal. This option is only applicable when Method is Indirect ELISA methods (IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA). It should be the same as the SecondaryAntibody for all samples or a subset of samples to make the blank valid.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Blank is not Null, automatically set to an stocked secondary antibody if BlankPrimaryAntibody is specified or the first SecondaryAntibody for the input samples.
Pattern Description: An object of type or subtype Model[Sample] or Object[Sample] or a prepared sample or Null.
Programmatic Pattern: ((ObjectP[{Model[Sample], Object[Sample]}] | _String) | Automatic) | Null
BlankSecondaryAntibodyDilutionFactor
The factor by which the BlankSecondaryAntibody concentration is reduced by SecondaryAntibodyDiluent, calculated as the total final volume consisting of both the BlankSecondaryAntibody and SecondaryAntibodyDiluent divided by the volume of the BlankSecondaryAntibody. When preparing the secondary antibody master mix in SecondaryAntibodyDilutionContainers, the total volume of prepared master mix is scaled across the input samples, standards and blanks, based on the number of samples that share the same SecondaryAntibody and SecondaryAntibodyDilutionFactor. For example, if BlankSecondaryAntibodyDilutionFactor is 1.25 and BlankSecondaryAntibodyVolume is 100 Microliters, the secondary antibody master mix is prepared by combining 100 Microliters of undiluted SecondaryAntibody with 25 Microliters of SecondaryAntibodyDiluent, assuming no duplicate dilutions are made. If one input sample shares the same SecondaryAntibody and its dilution factor with this blank, the master mix is prepared by mixing 200 Microliters of undiluted SecondaryAntibody with 50 Microliters of SecondaryAntibodyDiluent.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Blank is specified, automatically set to the same value as the option SecondaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 1 and less than or equal to 1000 or Null.
Programmatic Pattern: (RangeP[1, 1000] | Automatic) | Null
BlankSecondaryAntibodyVolume
The volume of undiluted BlankSecondaryAntibody added to SecondaryAntibodyDilutionContainers to prepare the diluted BlankSecondaryAntibody by mixing with SecondaryAntibodyDiluent.
Default Calculation: If BlankSecondaryAntibody is specified and BlankSecondaryAntibodyDilutionFactor is not Null or 1, automatically set to 1.1*BlankSecondaryAntibodyImmunosorbentVolume/BlankSecondaryAntibodyDilutionFactor.
Pattern Description: Greater than or equal to 0 microliters and less than or equal to 200 milliliters or Null.
Programmatic Pattern: (RangeP[0*Microliter, $MaxRoboticTransferVolume] | Automatic) | Null
BlankSecondaryAntibodyImmunosorbentVolume
The volume of the BlankSecondaryAntibody (either diluted or undiluted) to be loaded on the ELISAPlate(s) for the immunosorbent step.
Default Calculation: If Method is IndirectELISA, IndirectSandwichELISA, or IndirectCompetitiveELISA and Blank is specified, automatically set to the same value as the option SecondaryAntibodyImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankBlockingVolume
The amount of BlockingBuffer that is dispensed into the corresponding wells of the ELISAPlate(s) during blocking step.
Default Calculation: If Blocking is True and Blank is specified, automatically set to the same value as the option BlockingVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
BlankAntibodyComplexImmunosorbentVolume
The volume of the Blank-Antibody Complex to be loaded on the ELISAPlate(s). In DirectCompetitiveELISA and IndirectCompetitiveELISA, this step enables the free BlankPrimaryAntibody to bind to the BlankReferenceAntigen coated on the ELISAPlate(s). In FastELISA, this step enables the PrimaryAntibody-TargetAntigen-CaptureAntibody complex to bind to the BlankCoatingAntibody on the ELISAPlate(s).
Default Calculation: If the Method is set to DirectCompetitiveELISA, IndirectCompetitiveELISA, or FastELISA and Blank is specified, automatically set to the same value as the option SampleAntibodyComplexImmunosorbentVolume.
Pattern Description: Greater than or equal to 0.5 microliters and less than or equal to 300 microliters or Null.
Programmatic Pattern: (RangeP[$MinRoboticTransferVolume, Experiment`Private`$MaxWashVolume] | Automatic) | Null
Post Experiment
SamplesInStorageCondition
The non-default conditions under which the SamplesIn of this experiment should be stored after the protocol is completed. If left unset, SamplesIn will be stored according to their current StorageCondition.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (Alternatives[SampleStorageTypeP | Disposal]) | Null
Index Matches to: experiment samples
PreImmunosorbentLigandStorageCondition
The condition under which the unused portion of PreImmunosorbentLigand should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
SpikeStorageCondition
The condition under which the unused portion of spike should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
CoatingAntibodyStorageCondition
The condition under which the unused portion of CoatingAntibody should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
CaptureAntibodyStorageCondition
The condition under which the unused portion of Capture Antibody stock sample should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
ReferenceAntigenStorageCondition
The condition under which the unused portion of ReferenceAntigen stock should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
PrimaryAntibodyStorageCondition
The condition under which the unused portion of PrimaryAntibody stock should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
SecondaryAntibodyStorageCondition
The condition under which the unused portion of SecondaryAntibody stock should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: experiment samples
StandardStorageCondition
The condition under which the unused portion of Standard stock should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
Index Matches to: Standard
BlankStorageCondition
The condition under which the unused portion of Blank should be stored.
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal or Null.
Programmatic Pattern: (SampleStorageTypeP | Disposal) | Null
ELISAPlateStorageCondition
The condition under which the ELISA plate(s) will be stored after the protocol is completed.
Default Value: Refrigerator
Pattern Description: {AmbientStorage, EnclosedAmbientStorage, Refrigerator, Freezer, DeepFreezer, CryogenicStorage, YeastIncubation, YeastShakingIncubation, BacterialIncubation, BacterialShakingIncubation, MammalianIncubation, ViralIncubation, CrystalIncubation, AcceleratedTesting, IntermediateTesting, LongTermTesting, UVVisLightTesting} or Disposal.
Programmatic Pattern: SampleStorageTypeP | Disposal
Model Input
PreparedModelContainer
Indicates the container in which a Model[Sample] specified as input to the experiment function will be prepared.
Default Calculation: If PreparedModelAmount is set to All and when the input model has a product associated with both Amount and DefaultContainerModel populated, automatically set to the DefaultContainerModel value in the product. Otherwise set to Model[Container, Vessel, "2mL Tube"].
Pattern Description: An object of type or subtype Model[Container] or Null.
Programmatic Pattern: (ObjectP[Model[Container]] | Automatic) | Null
Index Matches to: experiment samples
PreparedModelAmount
Indicates the amount of a Model[Sample] specified as input to the experiment function that will be prepared in the PreparedModelContainer. When set to All and the input model sample is not preparable, the entire amount of the input model sample that comes from one of the Products is prepared. The selected product must have both Amount and DefaultContainerModel populated, and it must not be a KitProduct. When set to All and the input model is preparable such as water, 1 Milliliter of the input model sample is prepared.
Default Calculation: Automatically set to 1 Milliliter.
Pattern Description: All or Count or Count or Mass or Volume or Null.
Programmatic Pattern: ((RangeP[1*Microliter, 20*Liter] | RangeP[1*Milligram, 20*Kilogram] | GreaterP[0*Unit, 1*Unit] | GreaterP[0., 1.] | All) | Automatic) | Null
Index Matches to: experiment samples